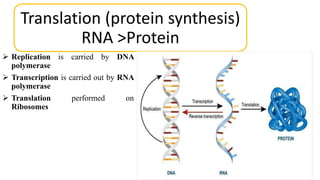
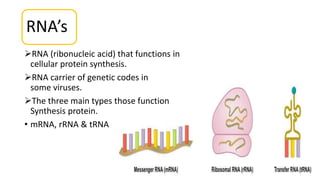
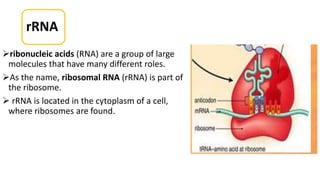
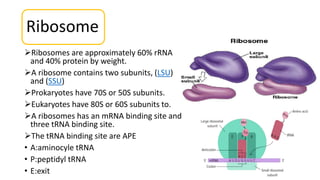
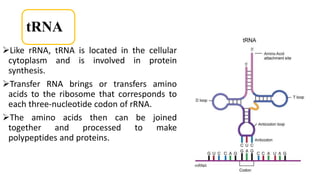
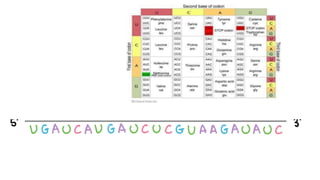
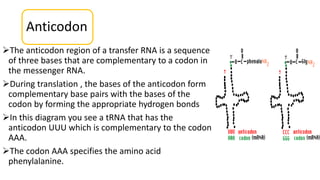
This document summarizes the process of protein synthesis through translation. It discusses how mRNA is transcribed from DNA and carries genetic codes to ribosomes. Ribosomes then translate mRNA into a polypeptide chain using transfer RNA (tRNA) and amino acids. The three phases of translation - initiation, elongation, and termination - are described in which mRNA codons are read and amino acids are linked together to form a protein.