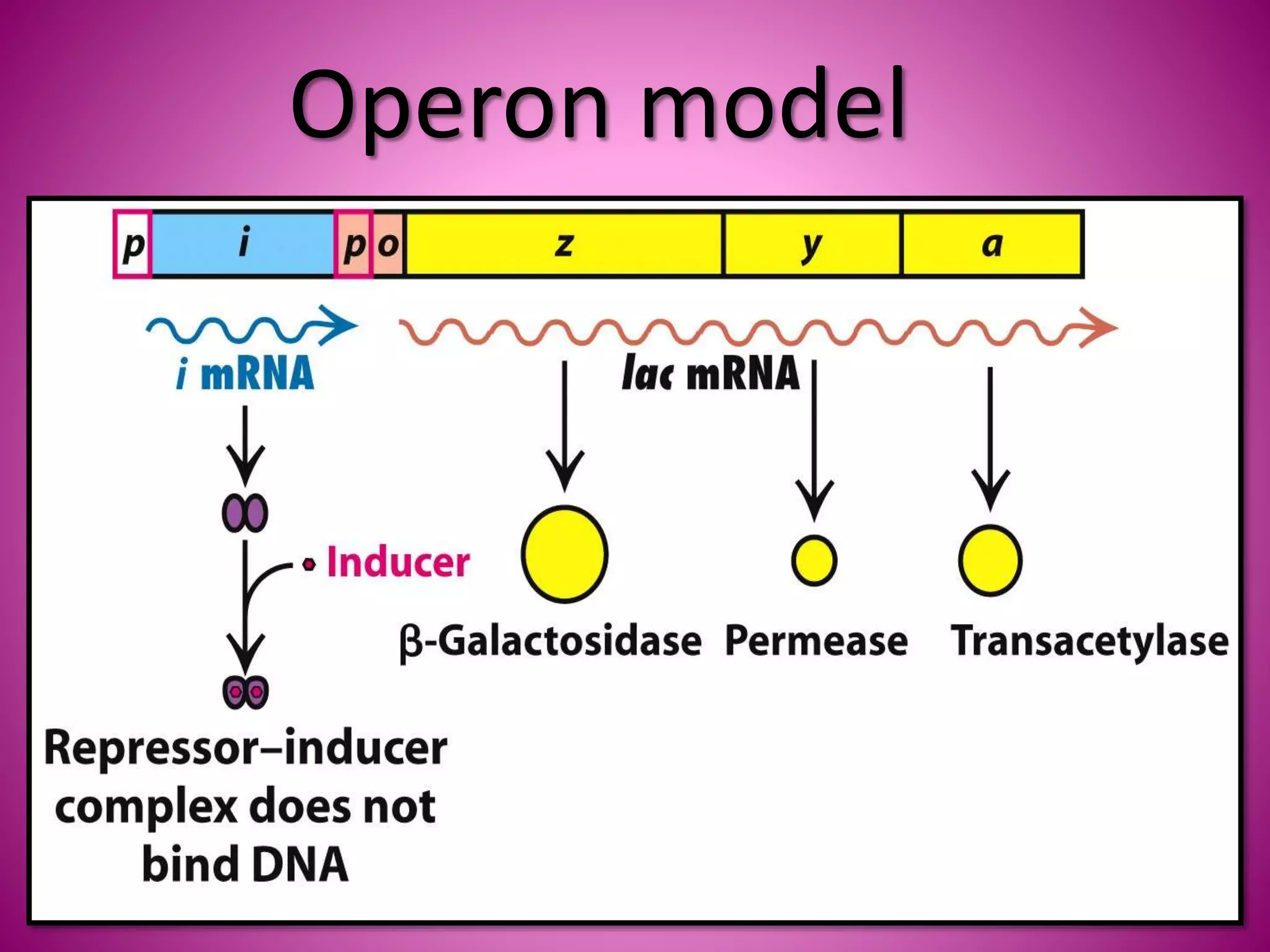
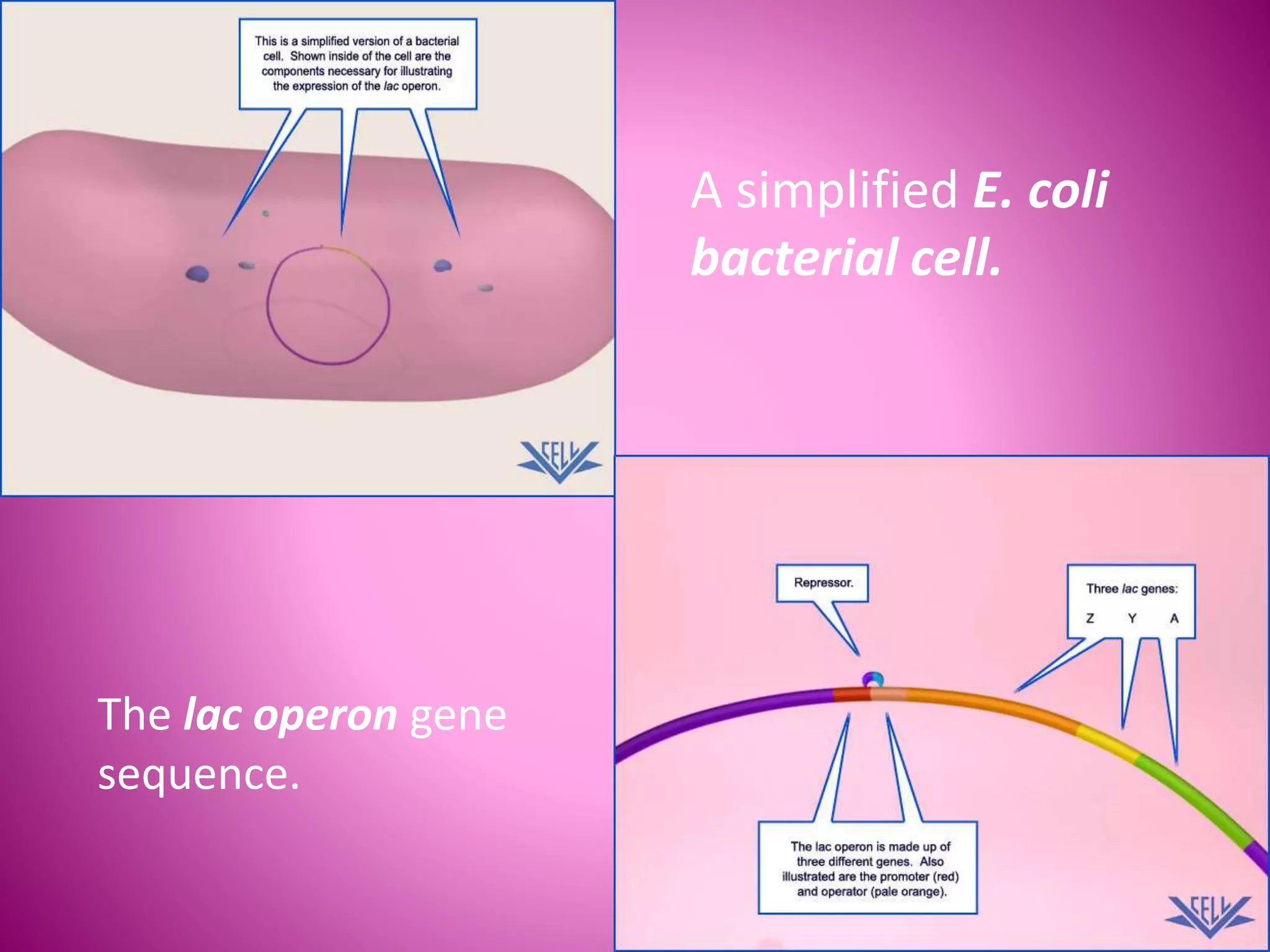
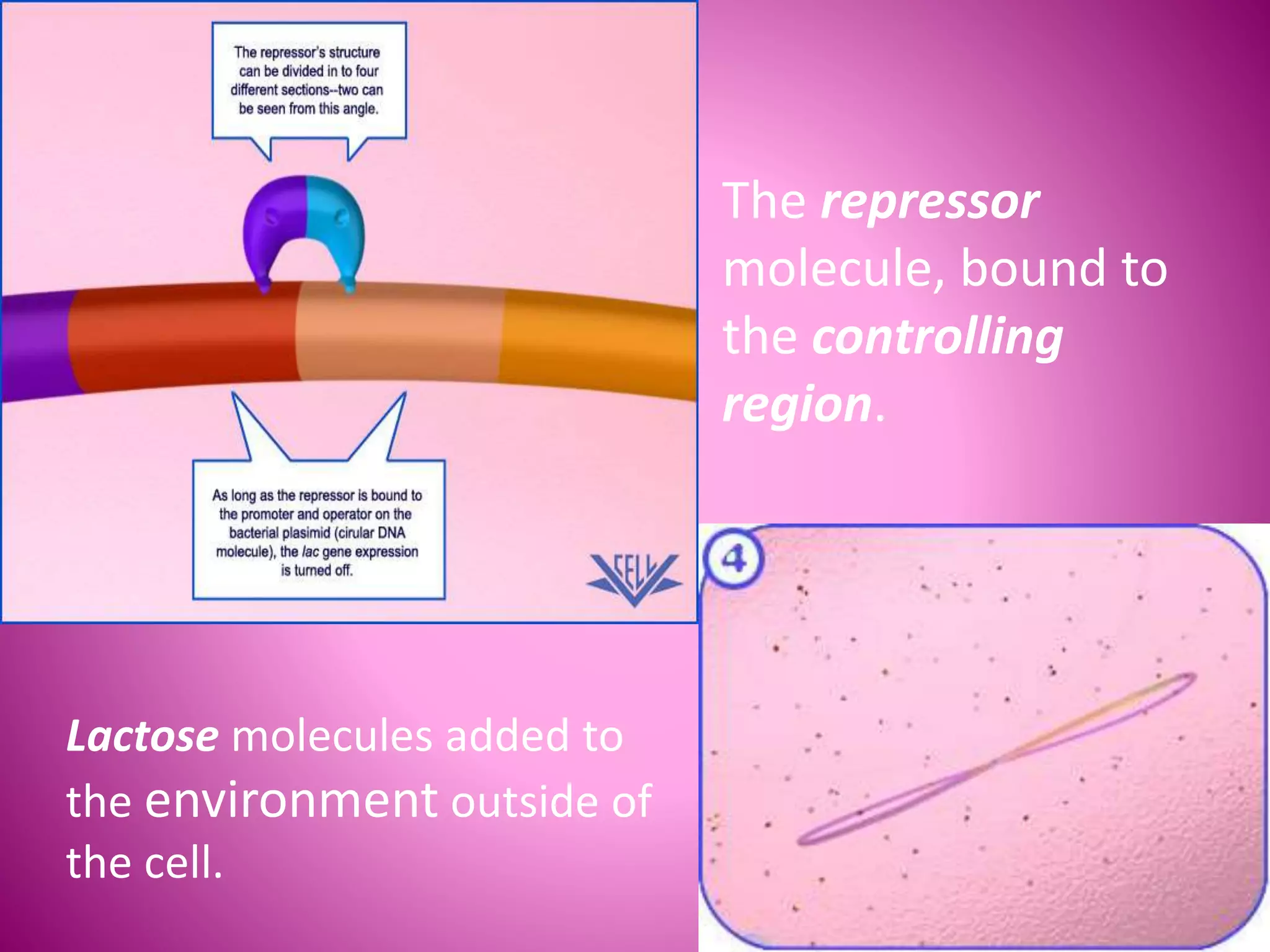
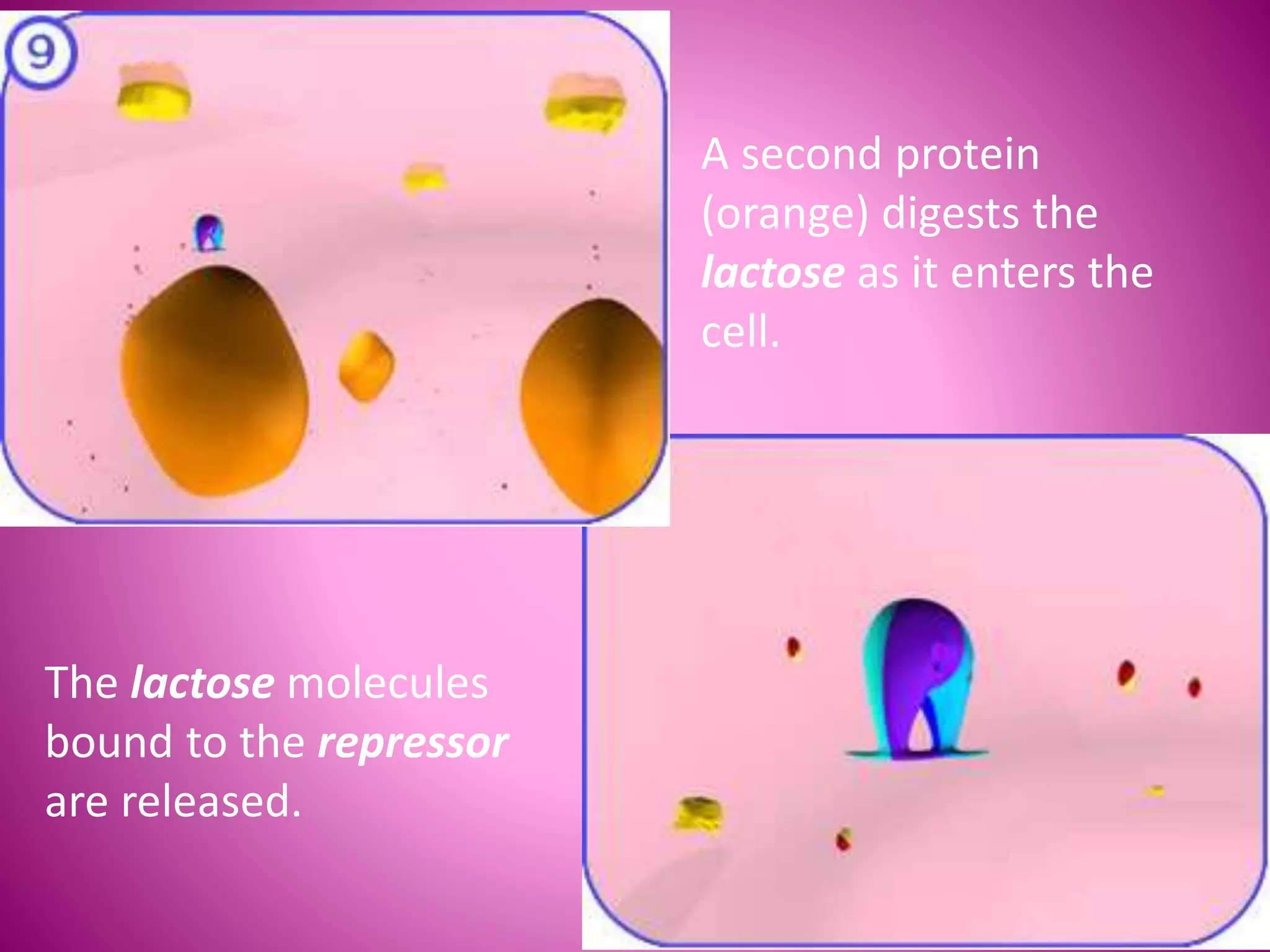
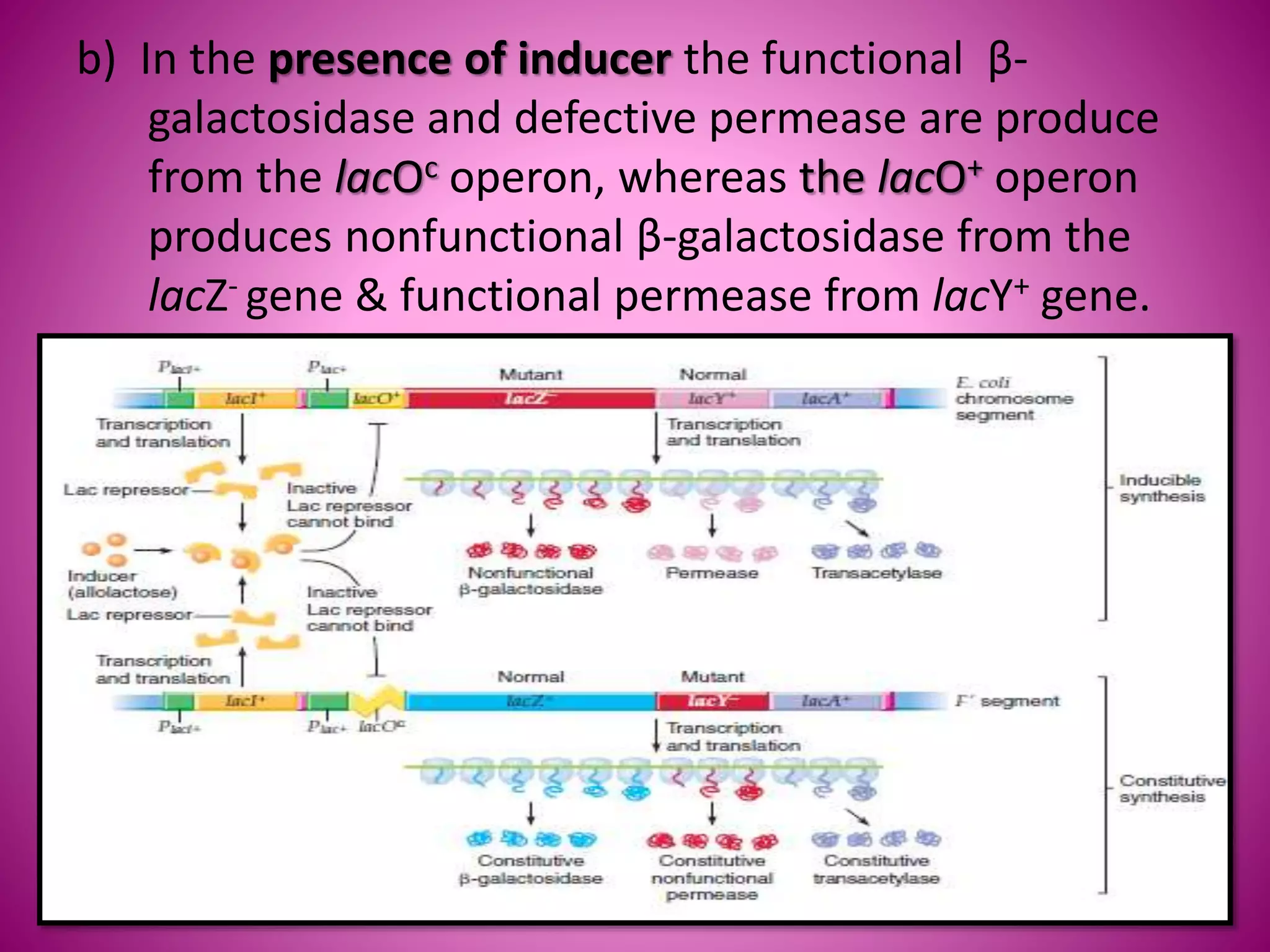
The lac operon controls the breakdown of lactose in E. coli bacteria. It consists of three structural genes (lacZ, lacY, lacA) that are regulated by a single promoter and operator region. In the absence of lactose, a lac repressor binds to the operator, preventing transcription of the structural genes. When lactose is present, it binds to the repressor and causes a conformational change, releasing it from the operator and allowing transcription. Mutations in the operator, structural genes, or promoter region provided insights into the operon's control mechanism.