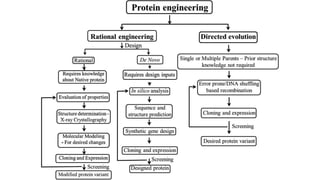
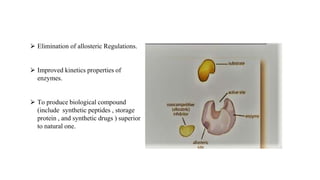
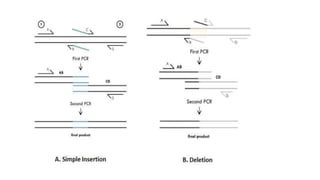
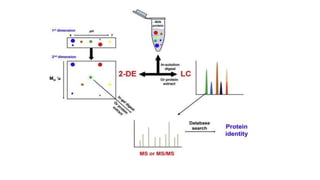
Protein engineering is a branch of biotechnology that modifies protein structure using genetic engineering techniques. The goals of protein engineering include developing proteins with useful functions for medicine, industry, and agriculture by making targeted changes to amino acids based on a protein's 3D structure. Common methods for protein engineering include site-directed mutagenesis and evolutionary methods involving random mutagenesis and selection. Proteome analysis studies all the proteins expressed by a genome at a given time using techniques like protein isolation, separation by SDS-PAGE or IEF, and identification through mass spectrometry.