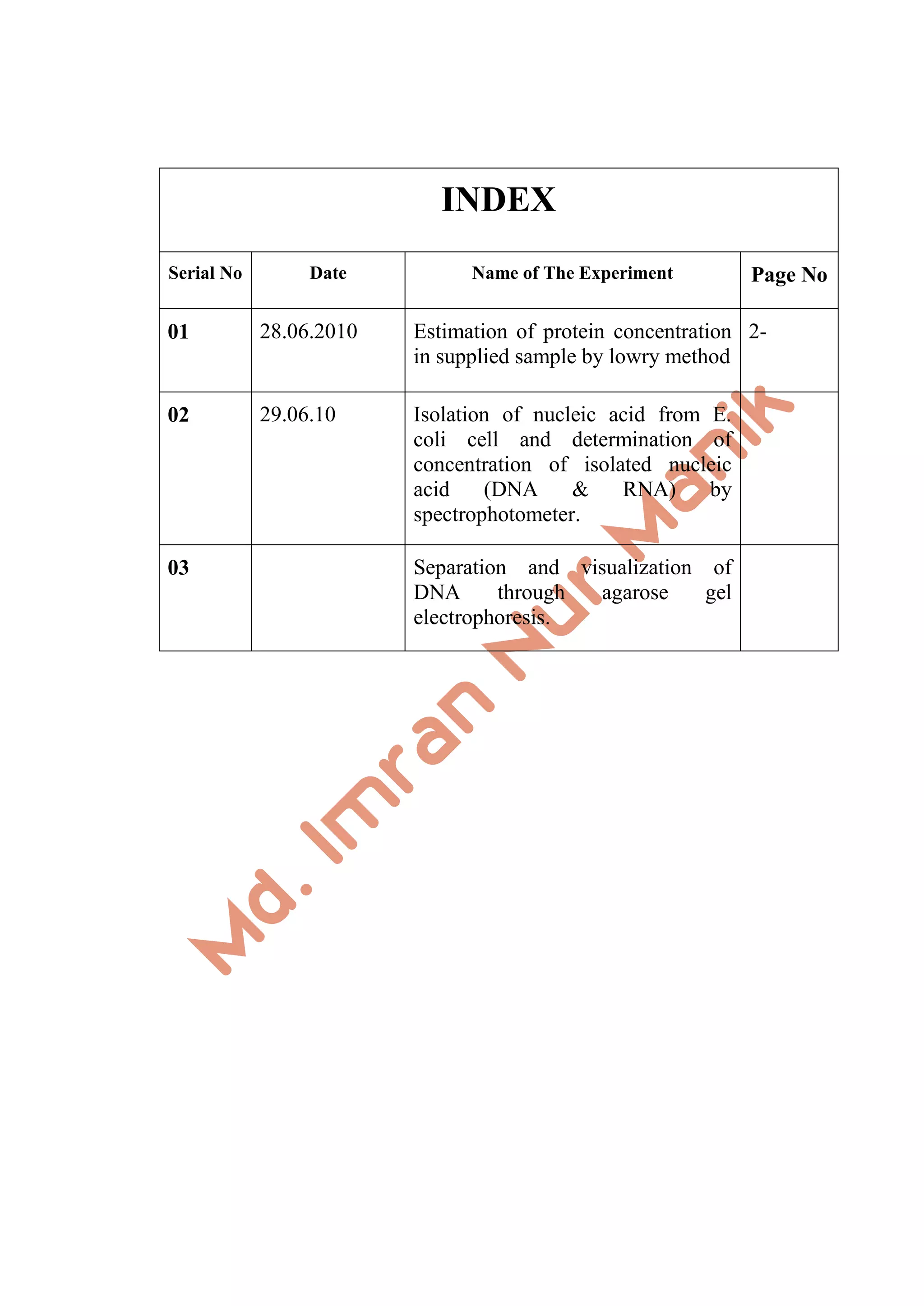
This document provides instructions for performing three experiments:
1) Estimating protein concentration using the Lowry method.
2) Isolating nucleic acids from E. coli and measuring concentration using a spectrophotometer.
3) Separating and visualizing DNA using agarose gel electrophoresis.
Reagents, equipment, and step-by-step procedures are described for each experiment. Calculations are shown for determining protein concentration from absorbance measurements.