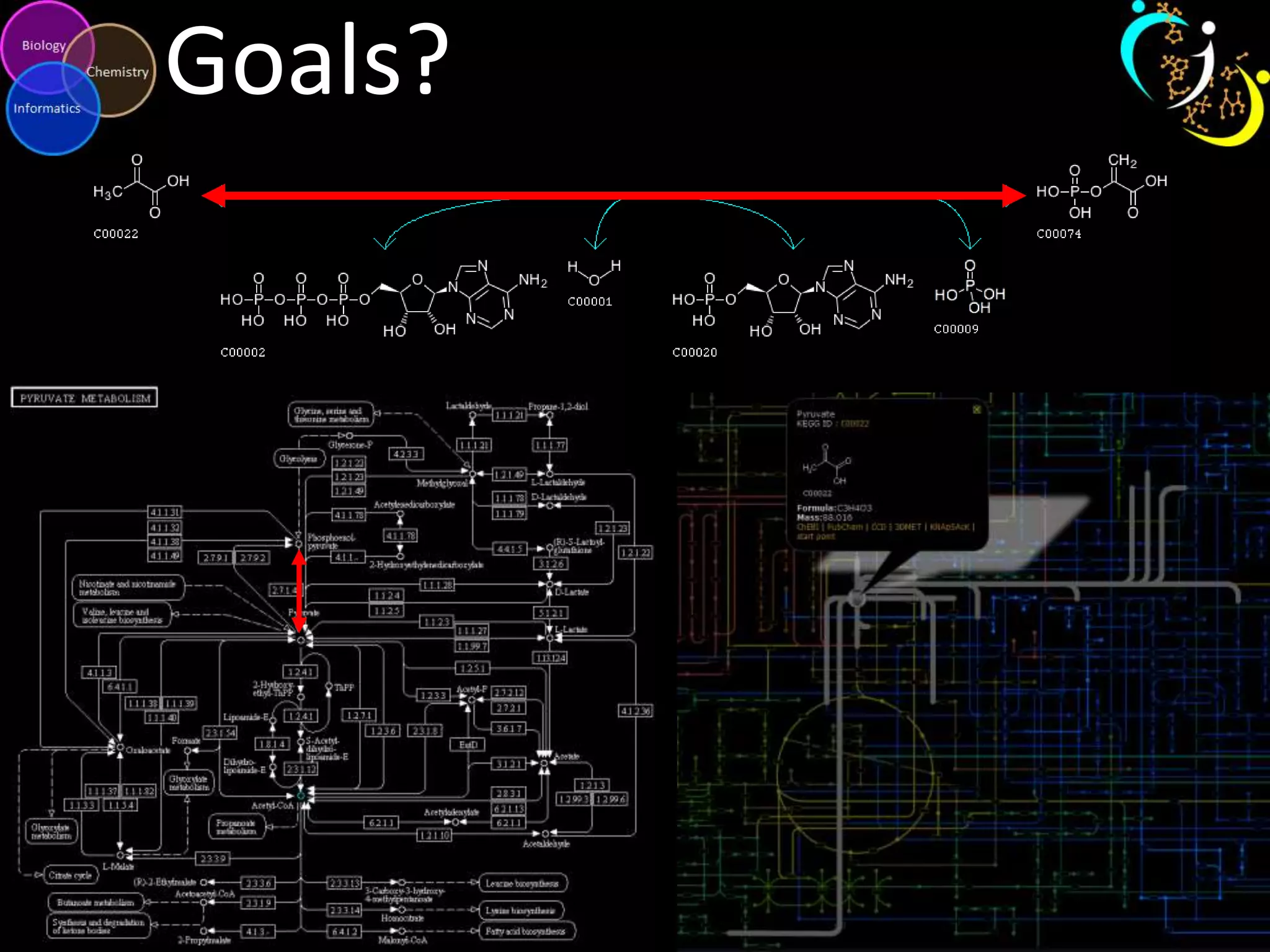
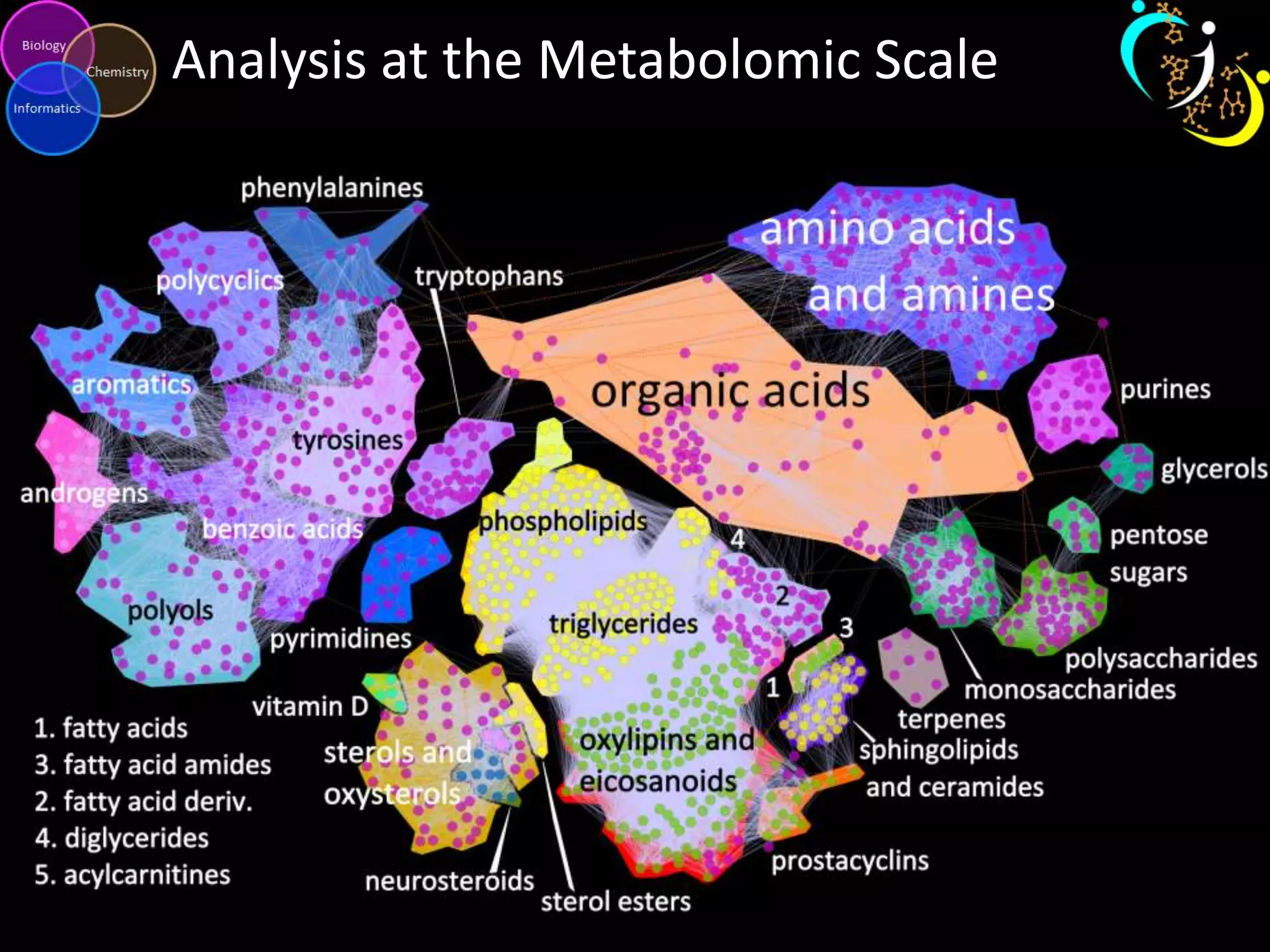
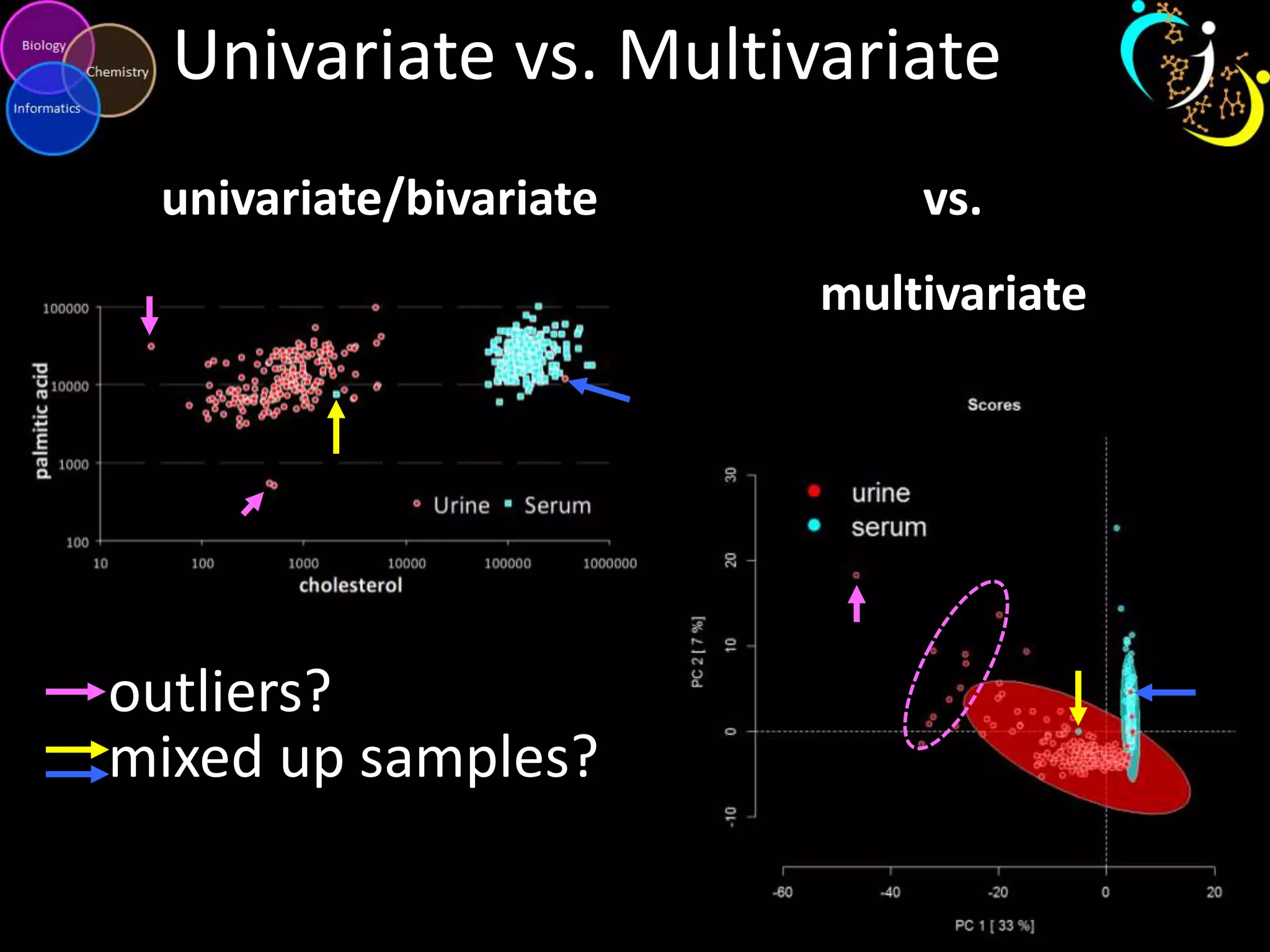
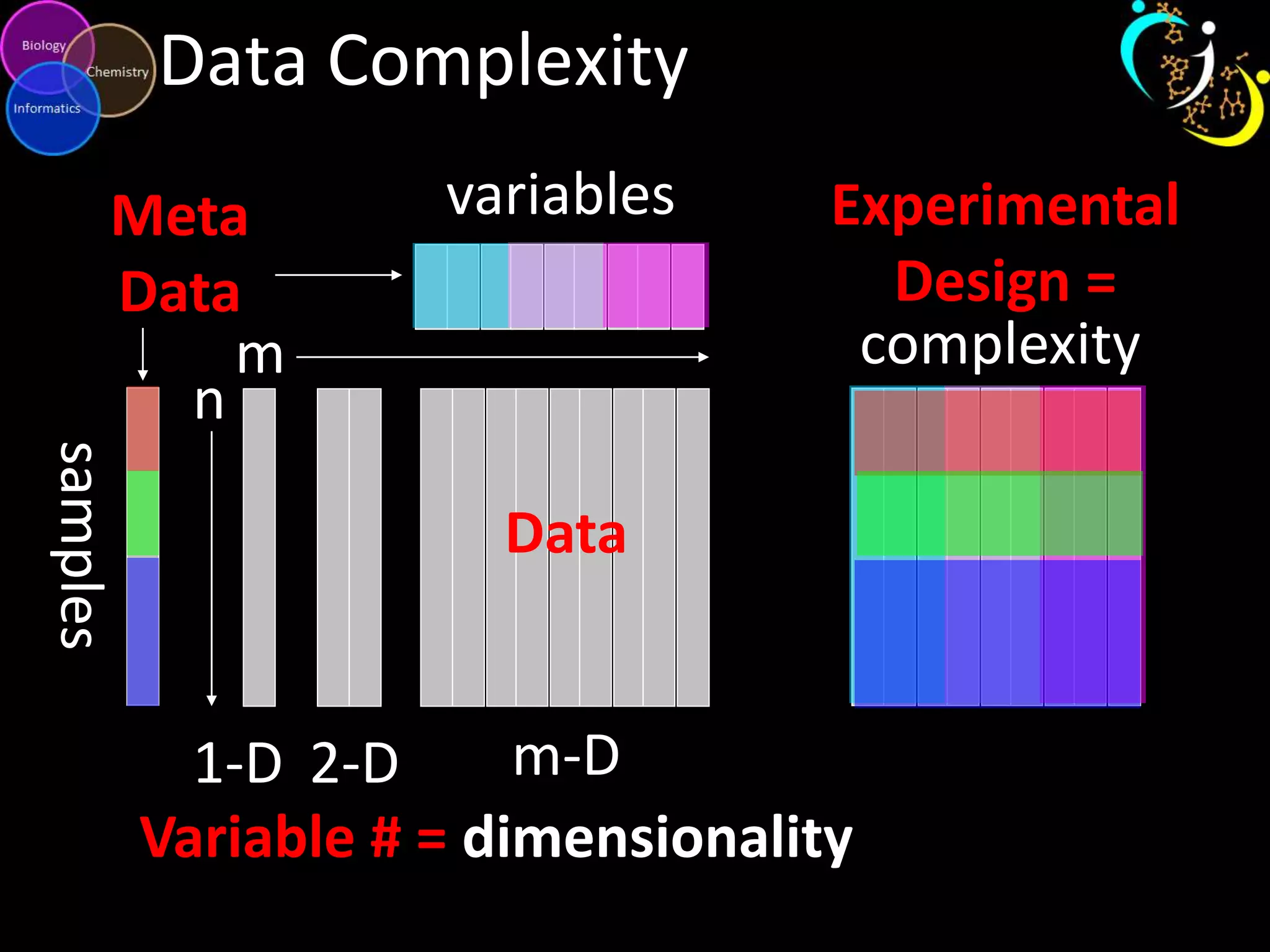
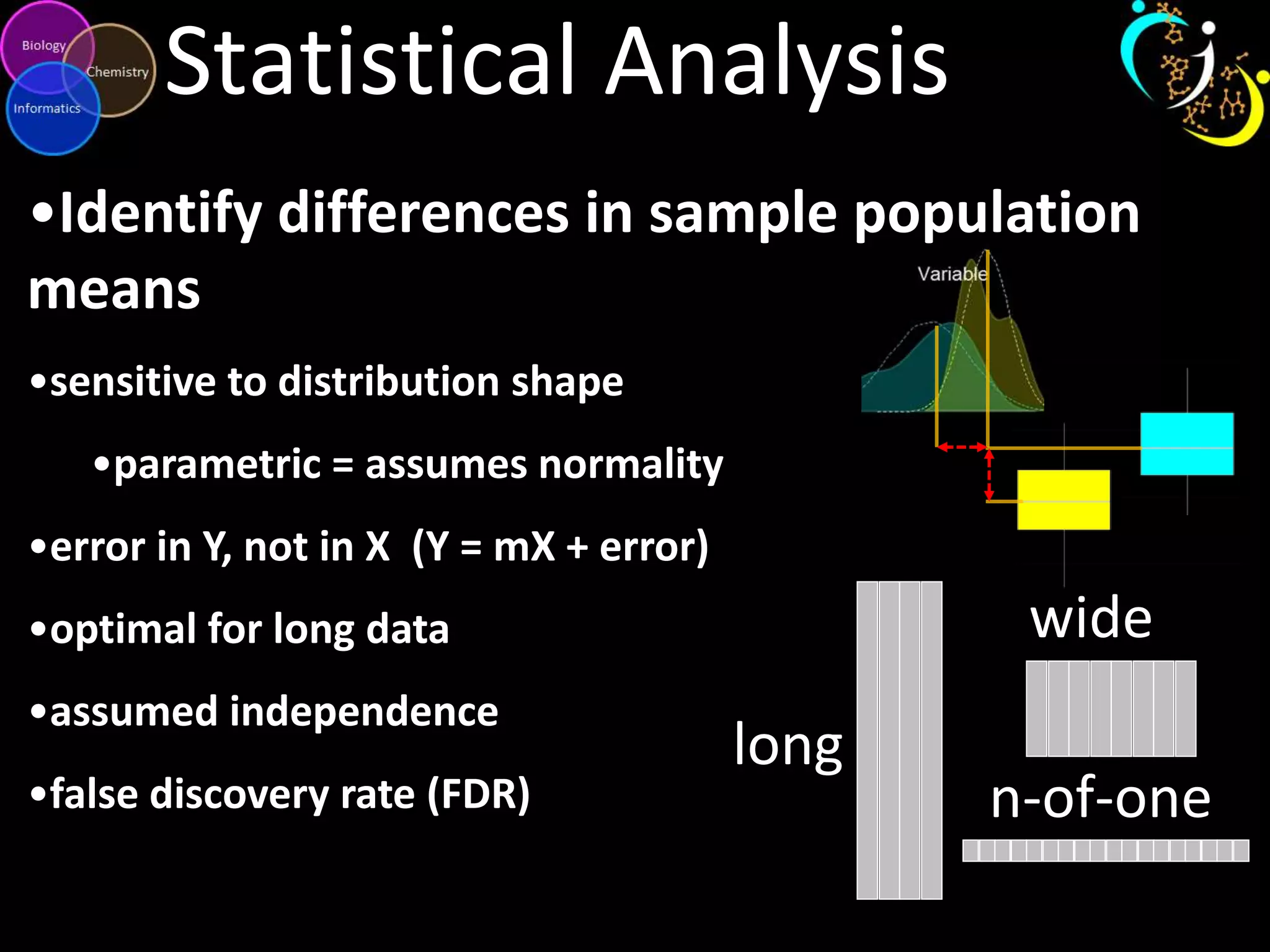
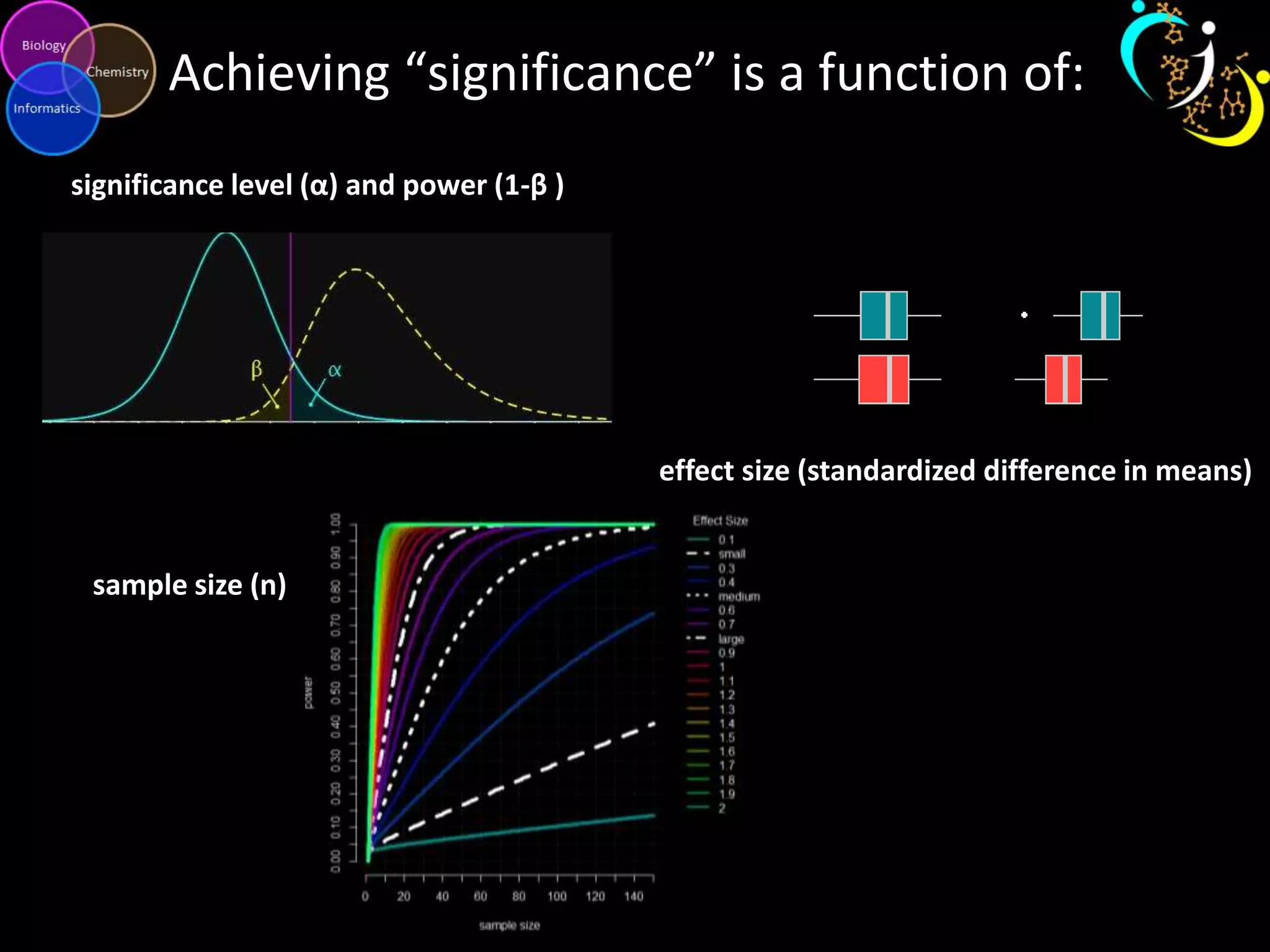
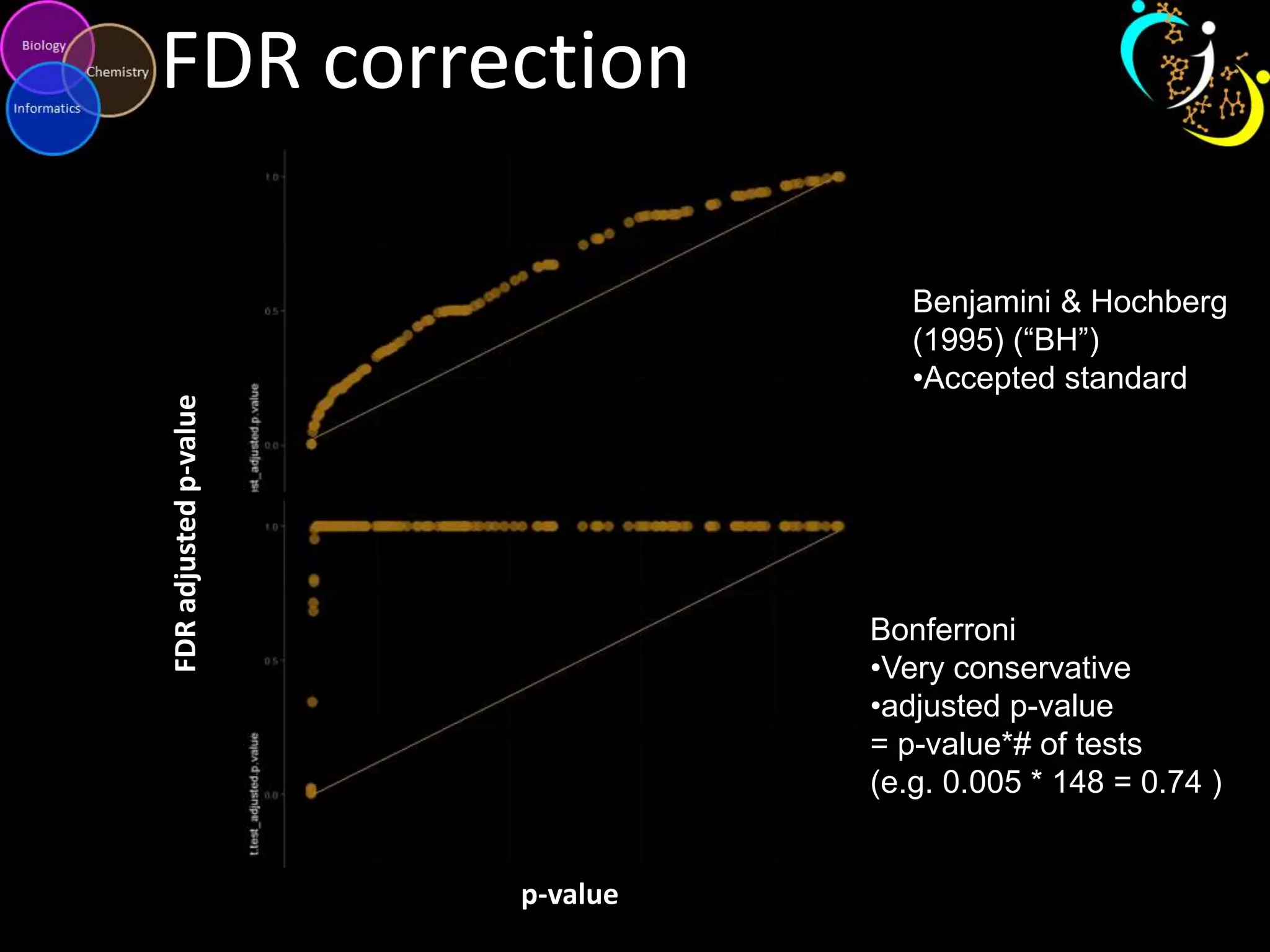
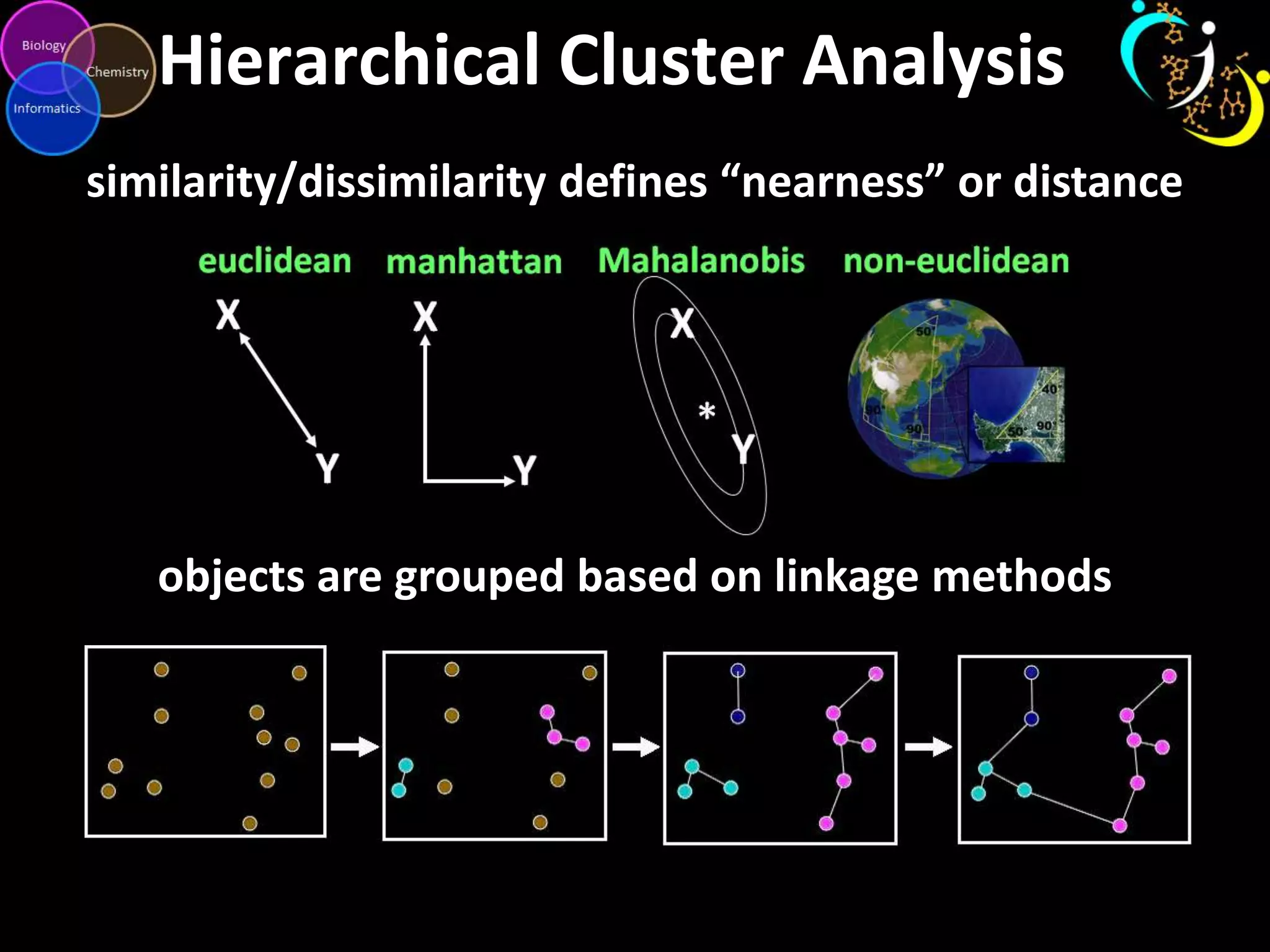
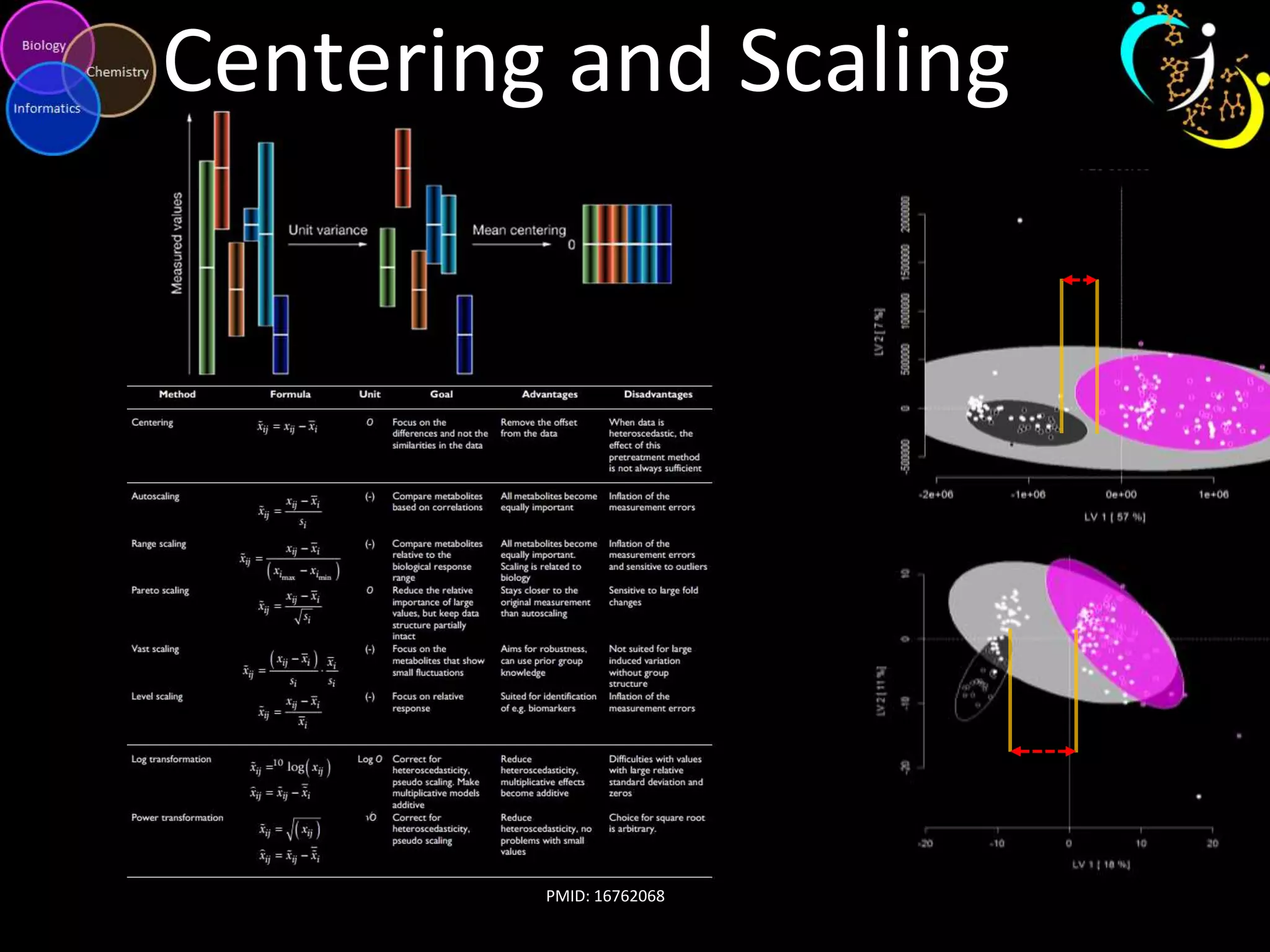
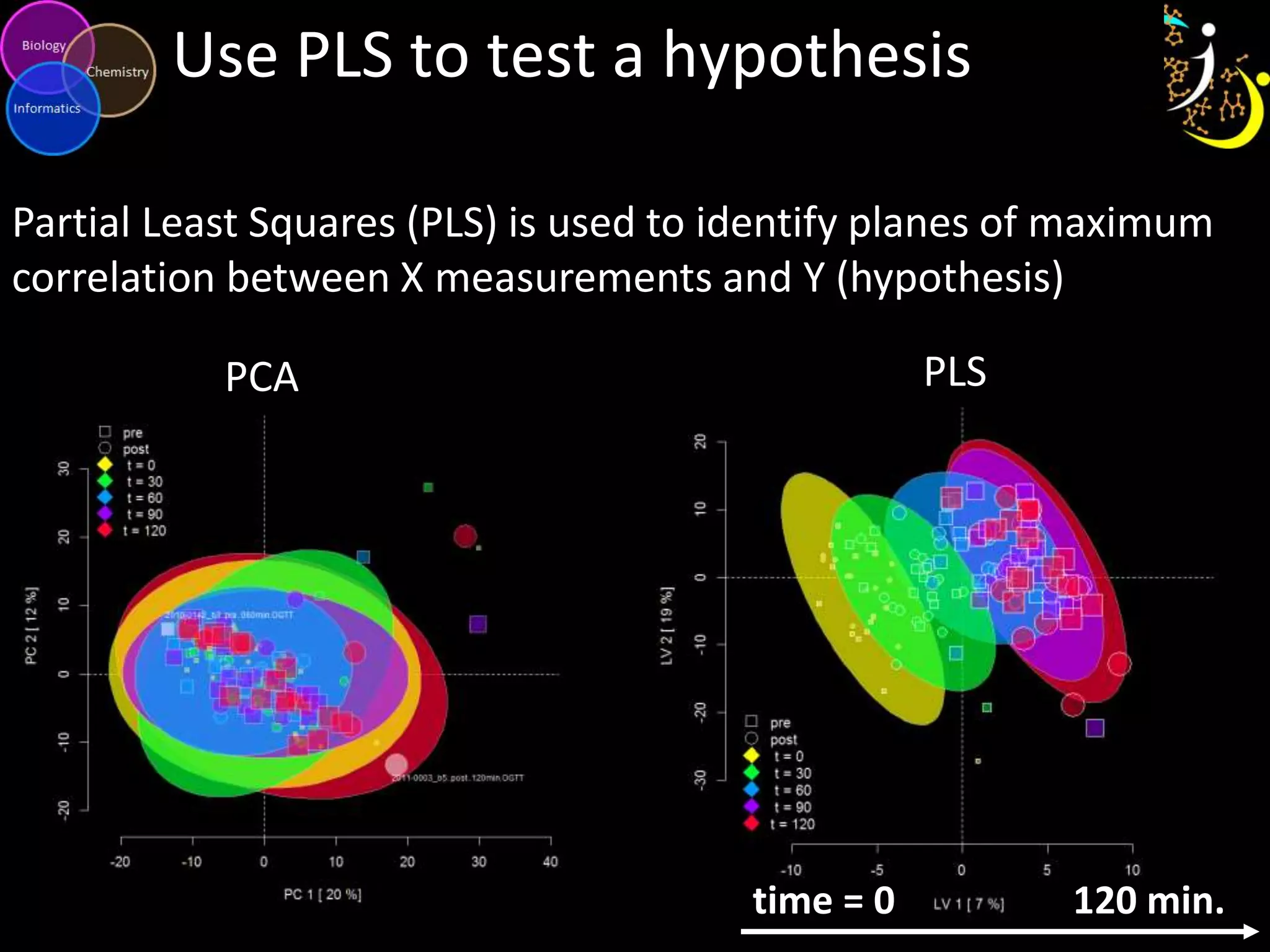
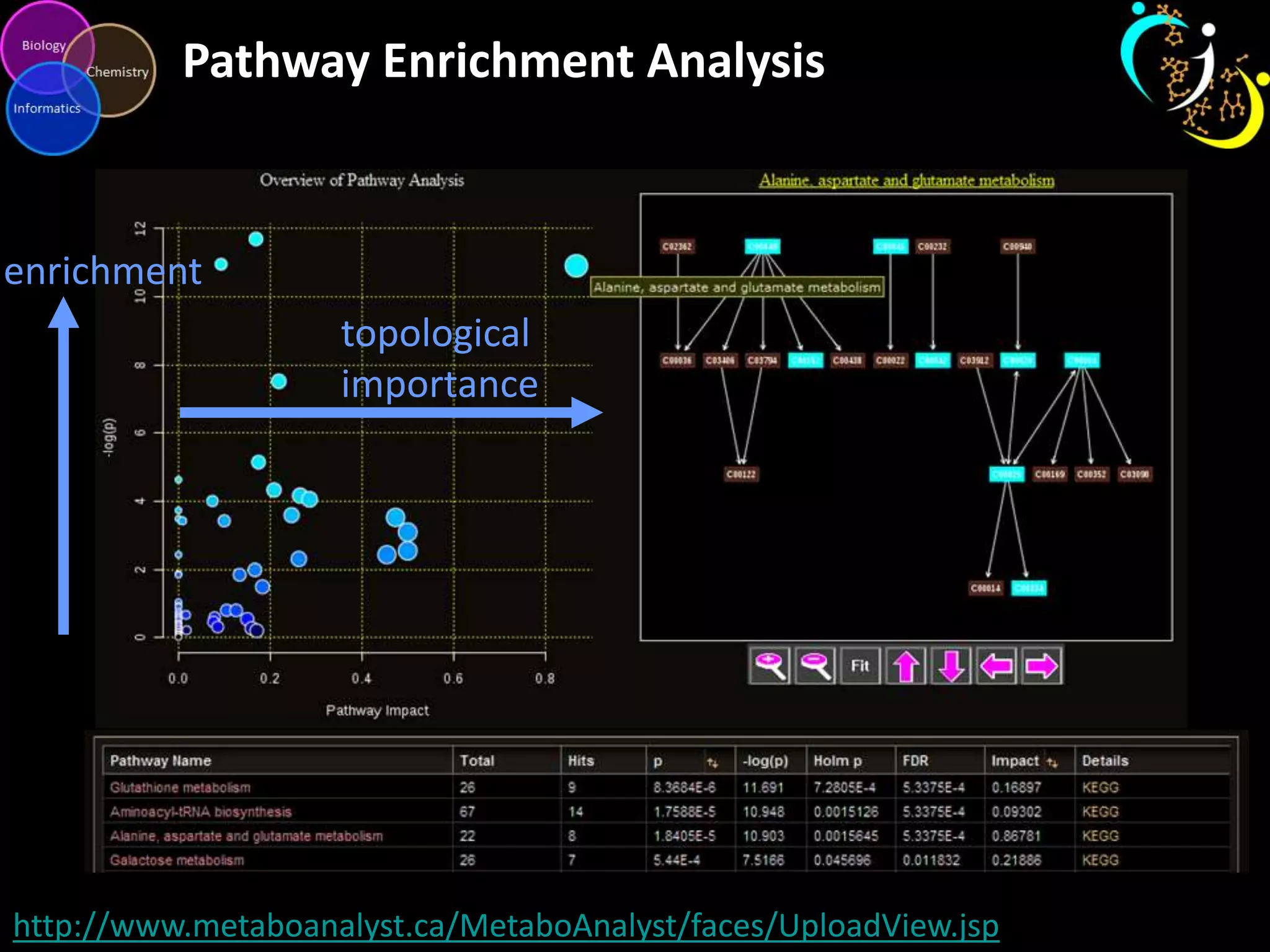
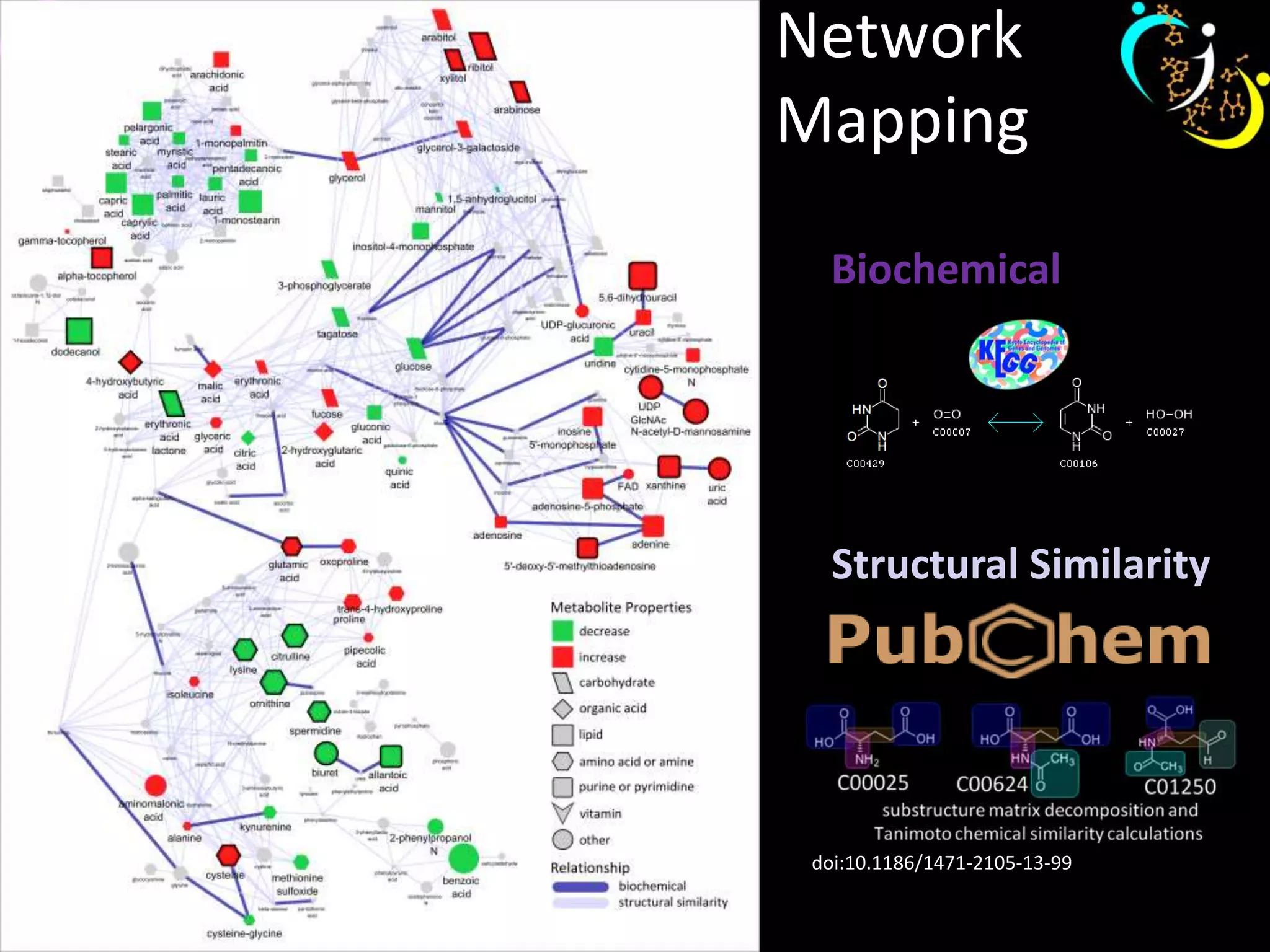
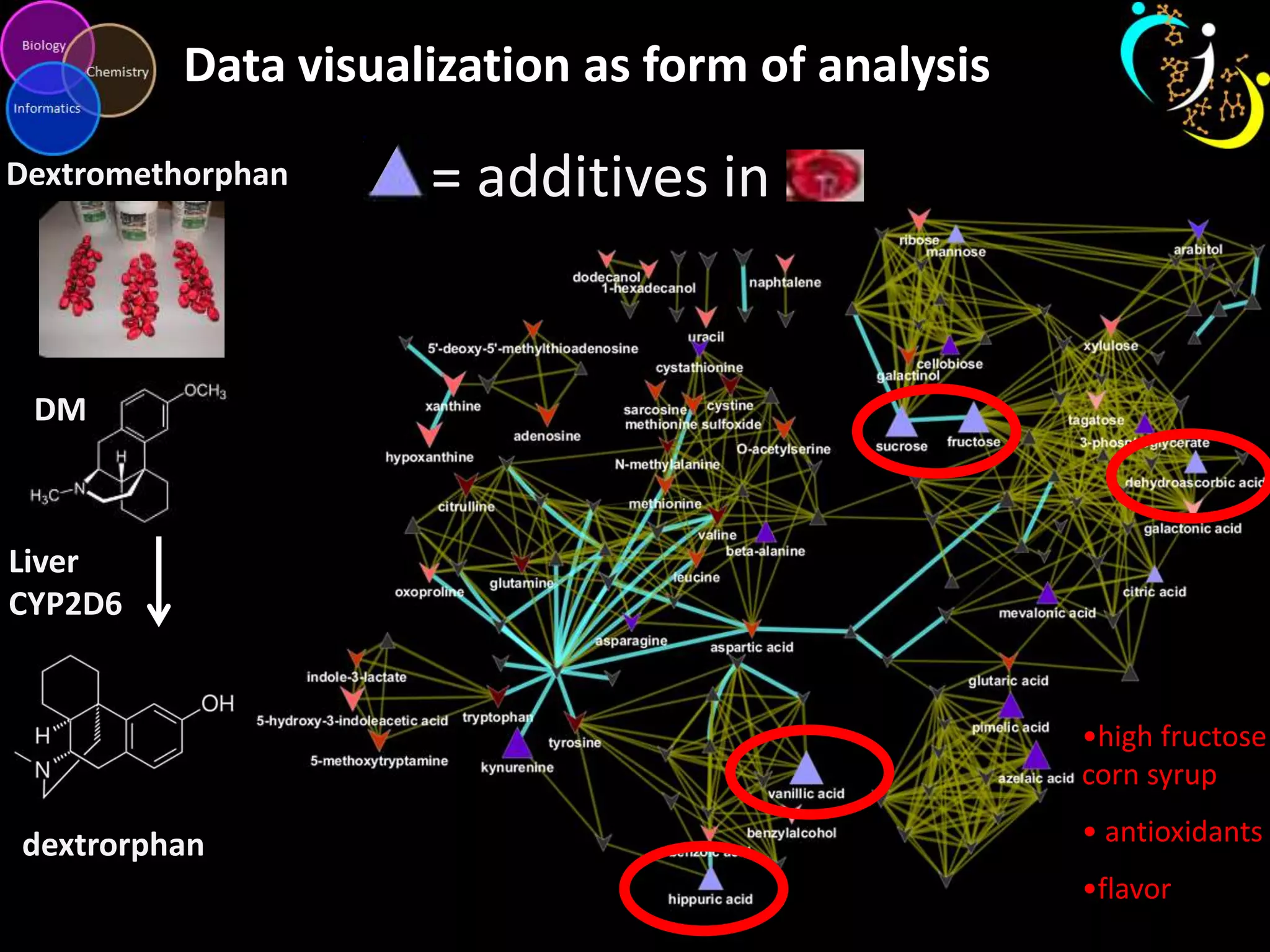
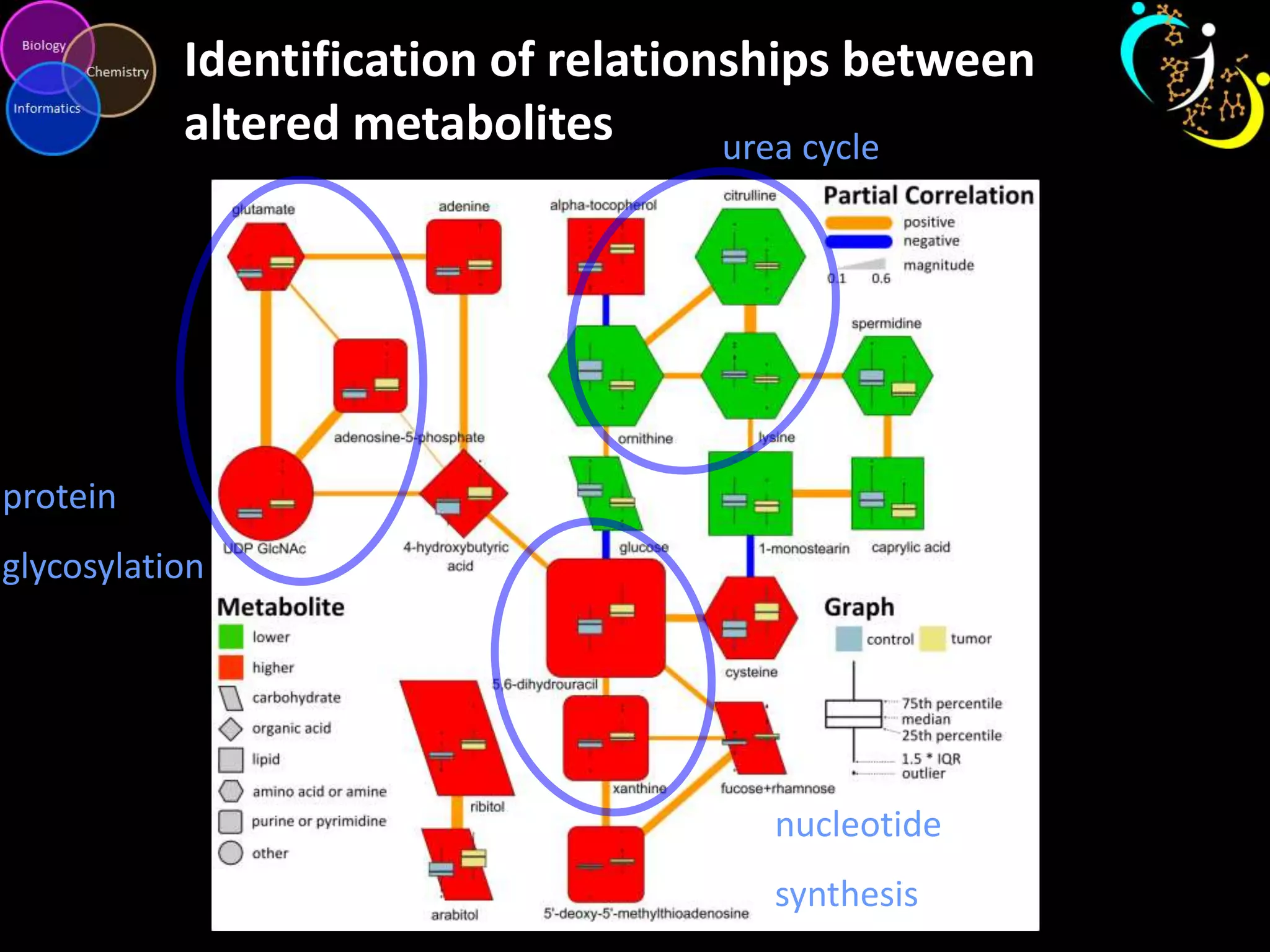
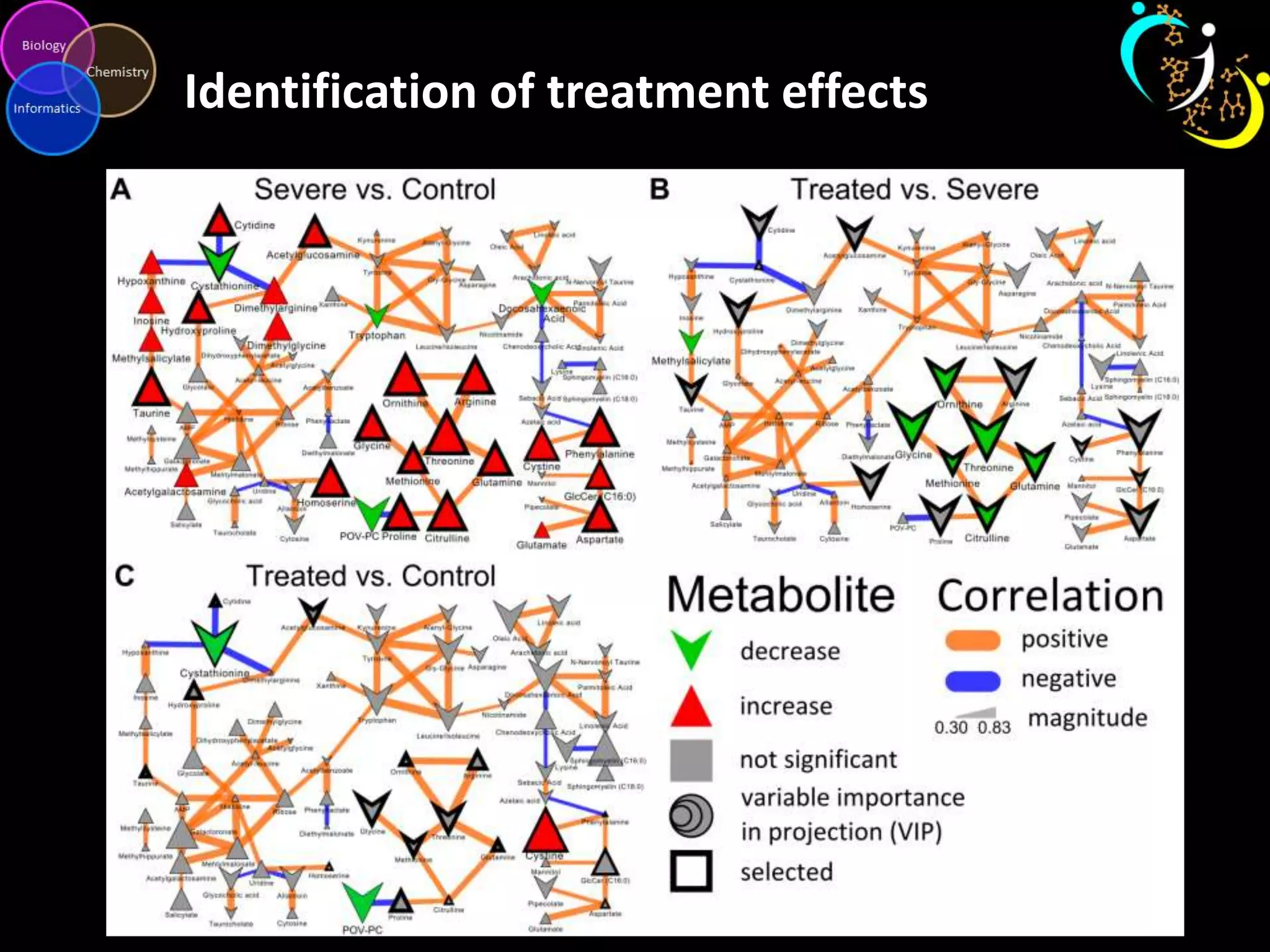
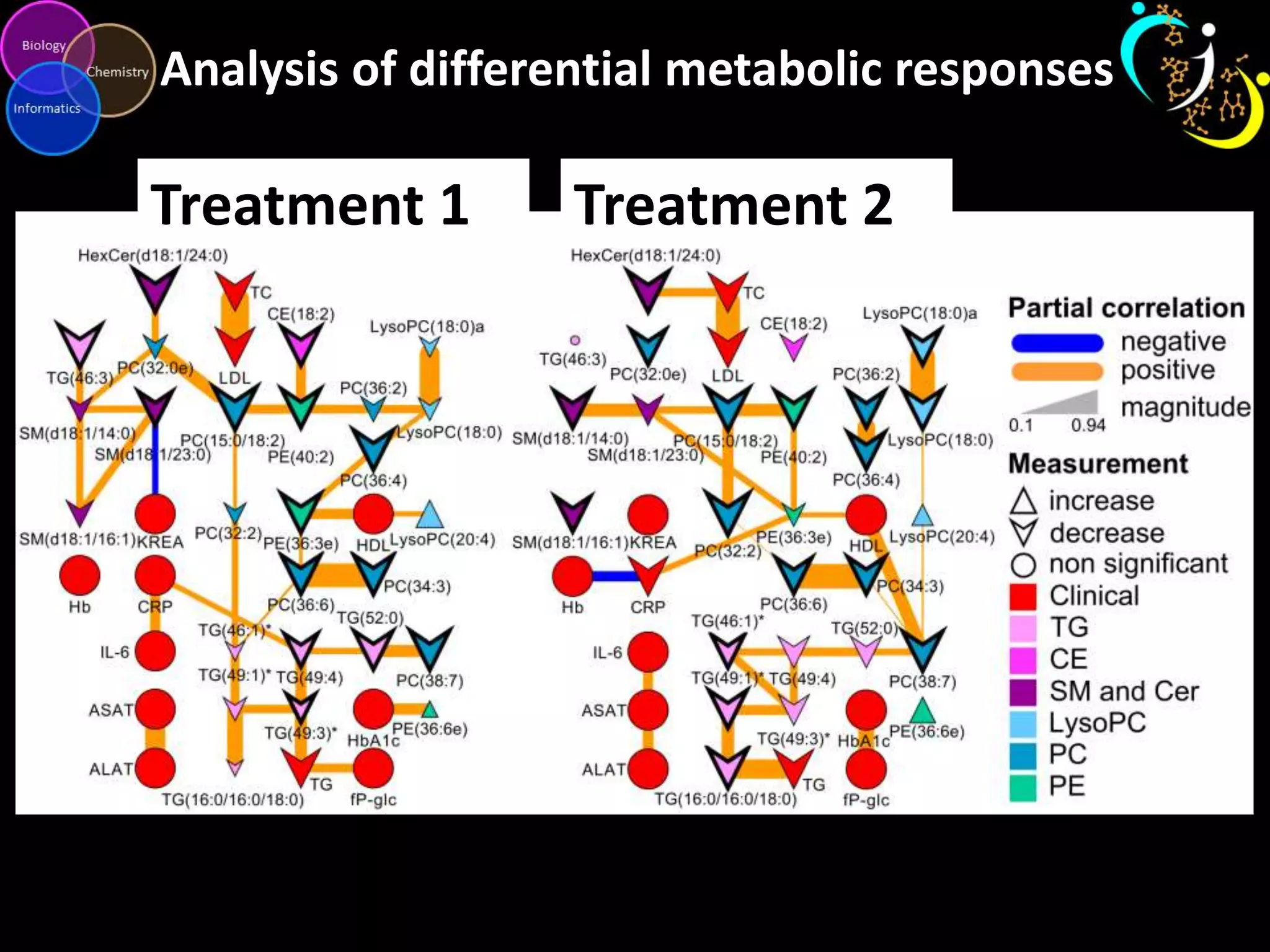
This document discusses metabolomic data analysis techniques for studying diseases. It analyzes over 13,000 biological samples per year using over 160,000 data points per study. Univariate and multivariate statistical analyses are described, with multivariate being preferred. Techniques include principal component analysis, partial least squares discriminant analysis, hierarchical clustering analysis, and pathway enrichment analysis. Visualization and network mapping tools are also discussed to identify relationships between altered metabolites and treatment effects.