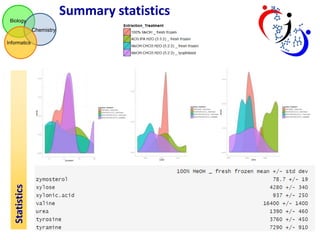
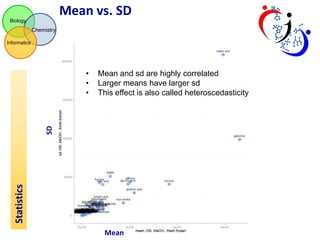
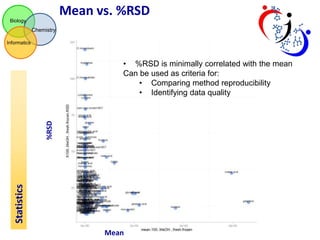
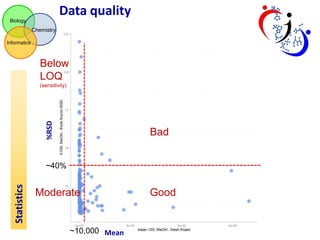
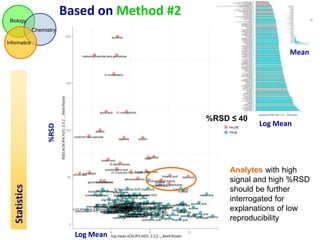
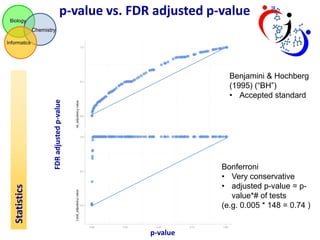
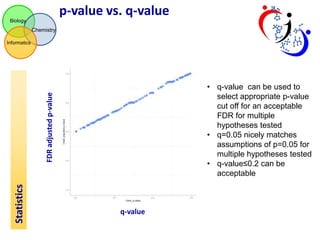
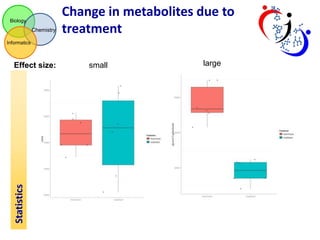
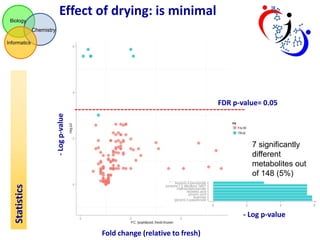
This document summarizes the analysis of different extraction and drying protocols for pumpkin leaf metabolites. It compares various metrics like mean, standard deviation, and percent relative standard deviation (%RSD) to identify the optimal sample processing method. Method #2, using acetonitrile, isopropanol and water in a 3:3:2 ratio, showed the lowest average %RSD and most metabolites below a 40% RSD cutoff, making it the optimal method. Statistical tests found minimal effect of drying on metabolite levels. A power analysis determined sample sizes needed to detect certain minimum effect sizes.