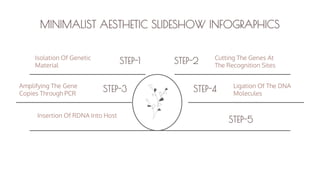
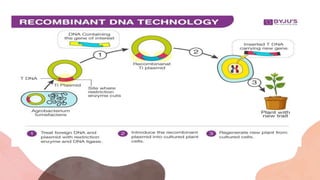
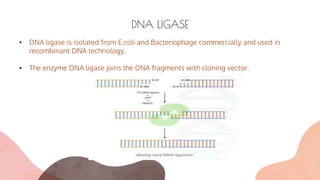
The document presents an introduction to recombinant DNA technology, a process that alters an organism's phenotype by integrating a genetically modified vector into its genome. It details the steps involved, including isolation, cutting, amplification, ligation, and insertion of DNA, as well as various enzymes crucial for these processes, such as DNA ligase and restriction endonucleases. The technology emerged from the discovery of restriction enzymes and is widely utilized in genetic engineering and biotechnology.







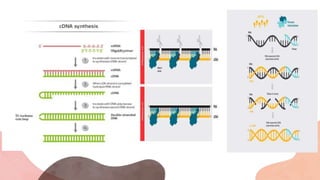
![RIBONUCLEASE –H [RNASE H]
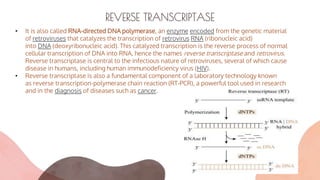
• Reverse transcriptases are enzymes composed of distinct domains that exhibit different
biochemical activities. RNA-dependent DNA polymerase activity and RNase H activity are the
predominant functions of reverse transcriptases, although depending on the source
organisms there are variations in functions, including, for example, DNA-dependent DNA
polymerase activity.
• RNase H cleaves the RNA template of the RNA:cDNA hybrid concurrently with
polymerization.The RNase H activity is undesirable for synthesis of long cDNAs because the
RNA template may be degraded before completion of full-length reverse transcription. The
RNase H activity may also lower reverse transcription efficiency, presumably due to its
competition with the polymerase activity of the enzyme.](https://image.slidesharecdn.com/introductiontordnatecchnology-231115042539-a2d3125f/85/Introduction-to-rDNA-Tecchnology-pptx-14-320.jpg)