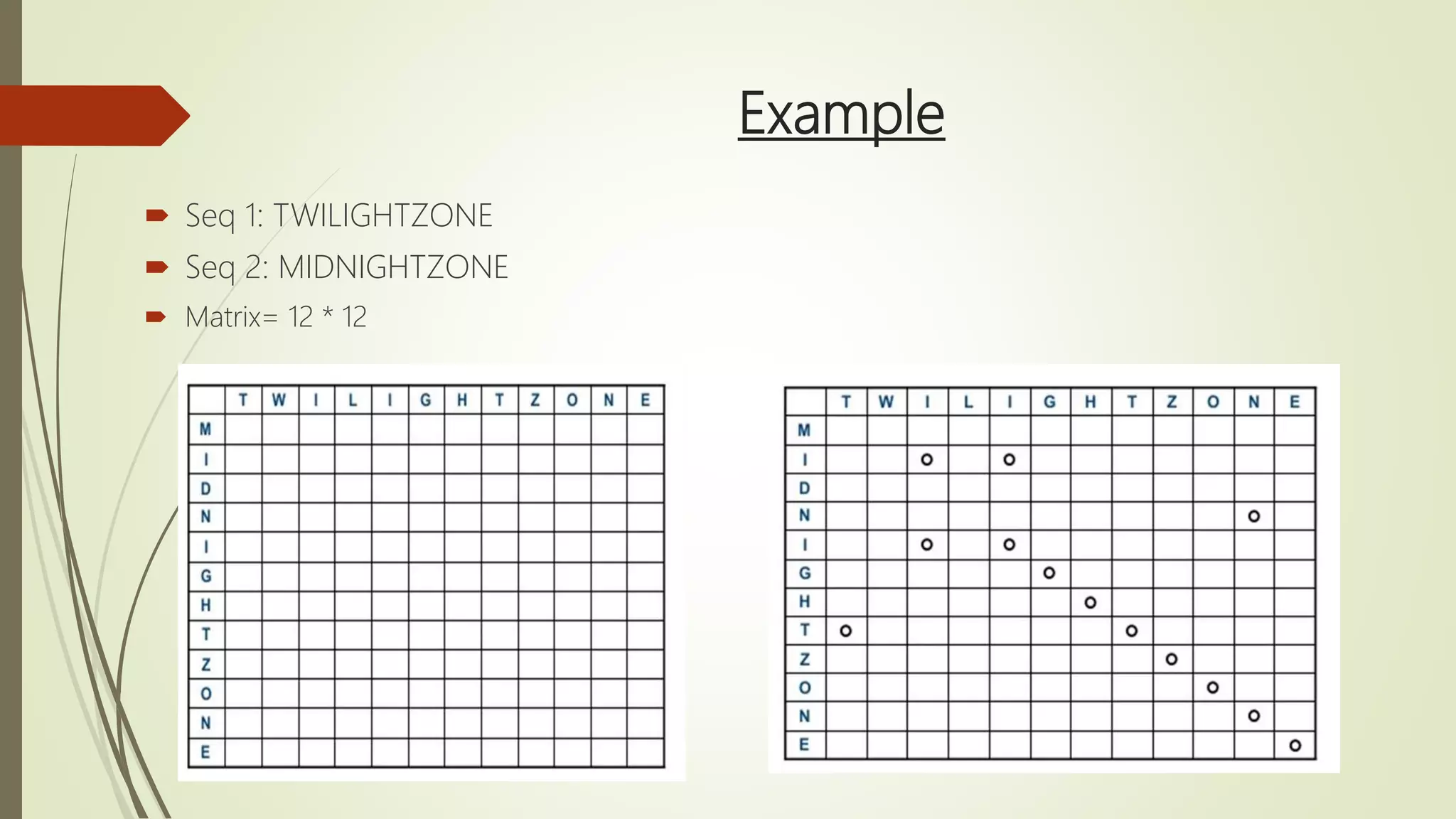
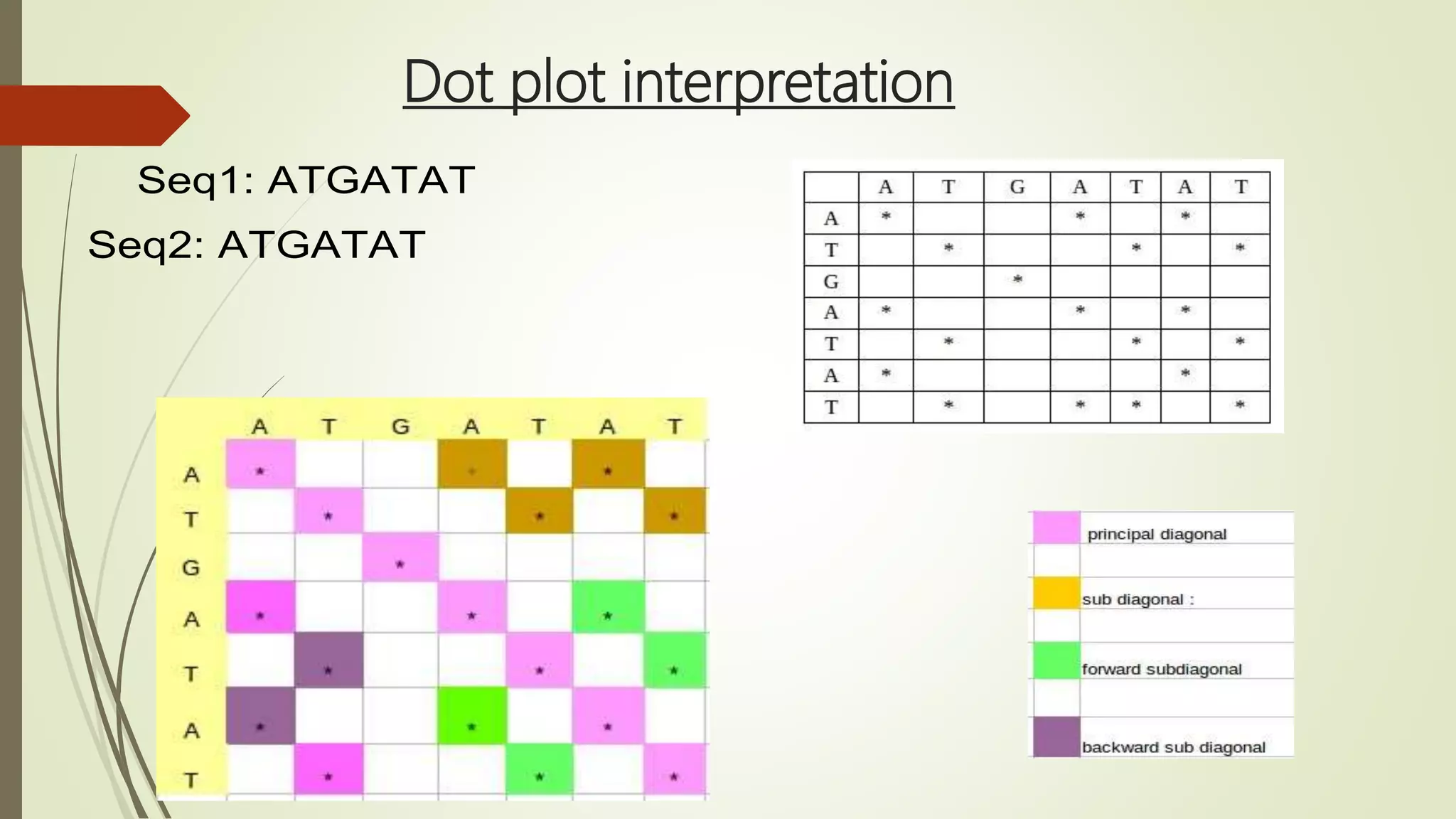
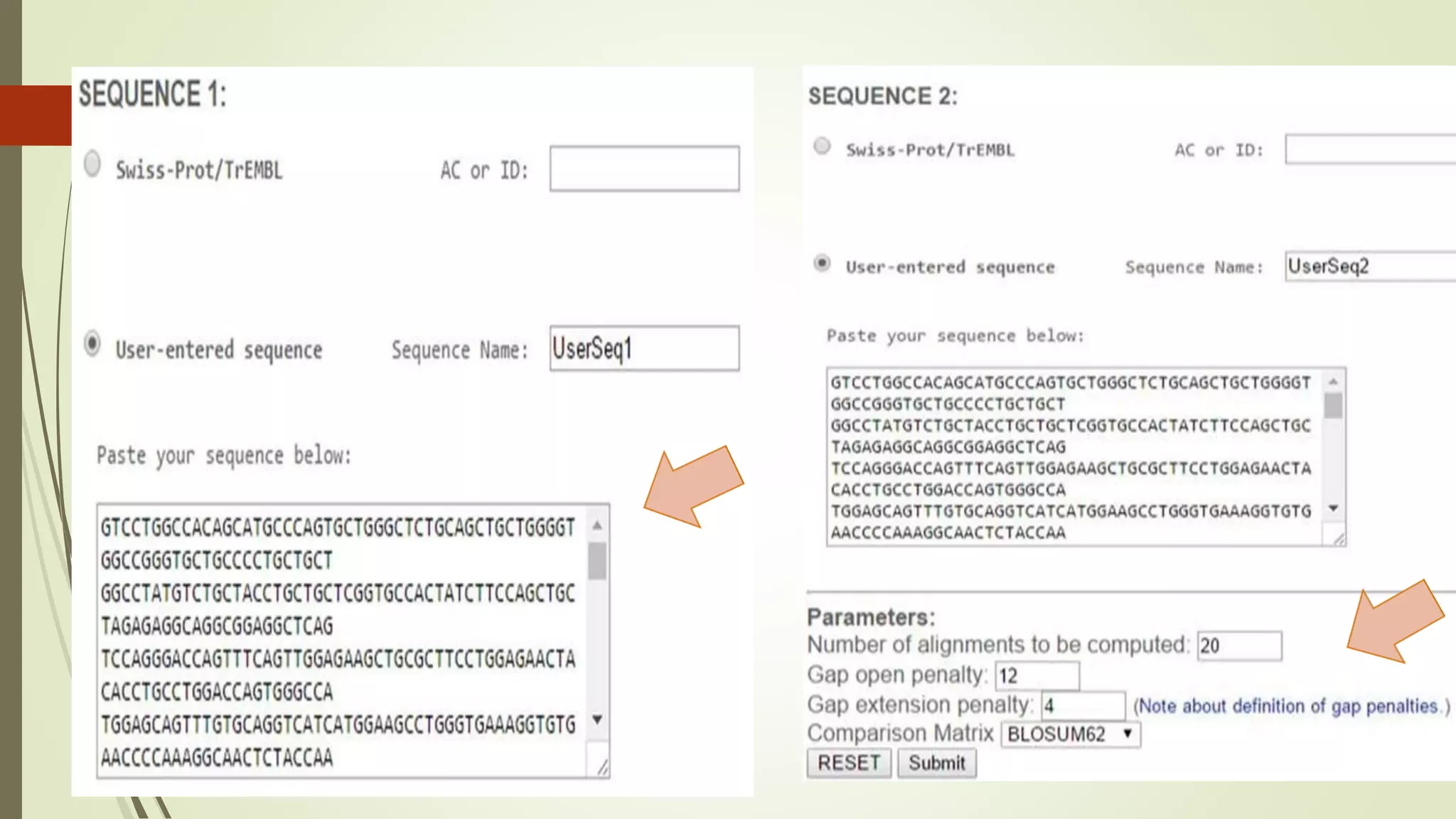
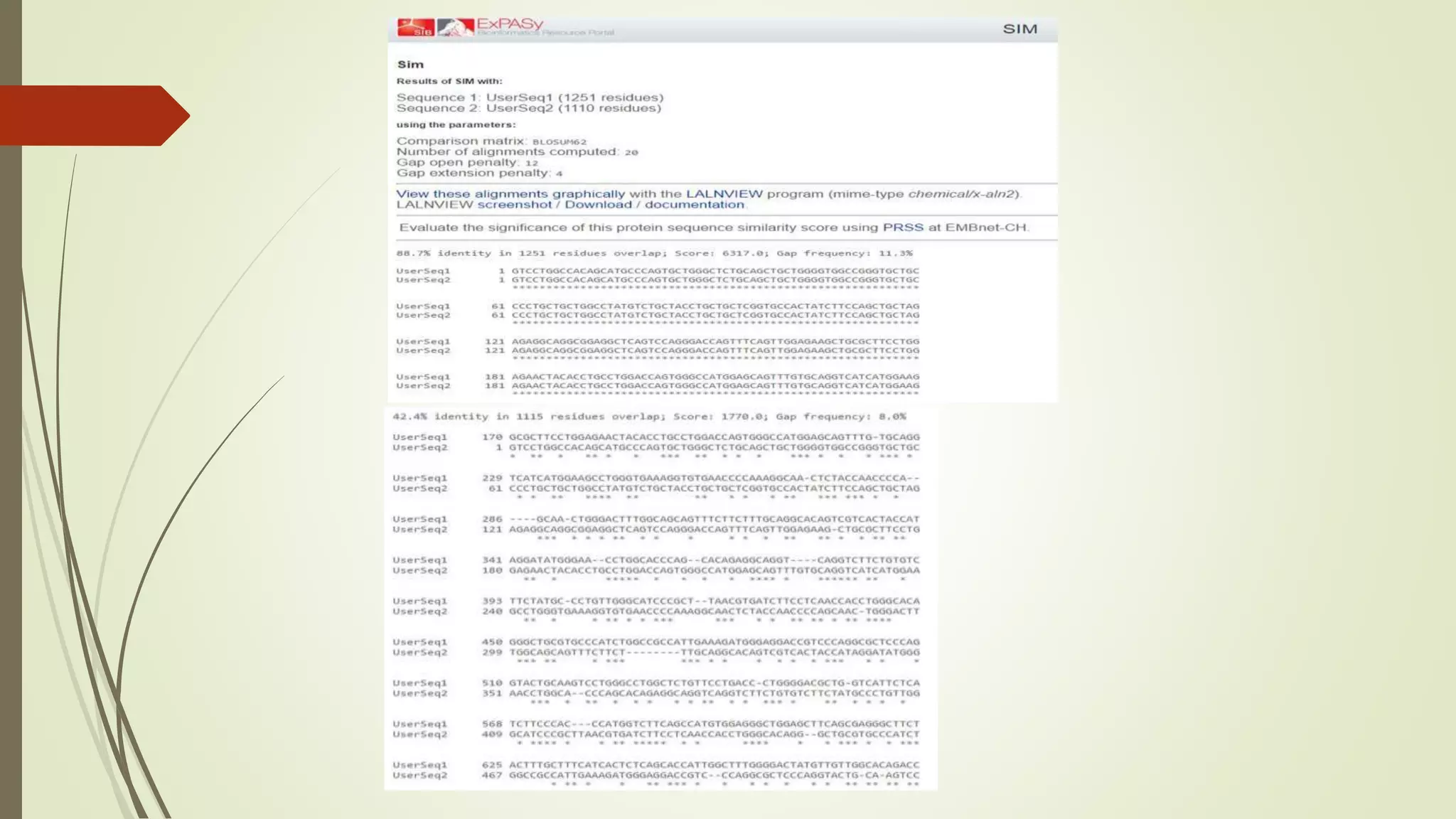
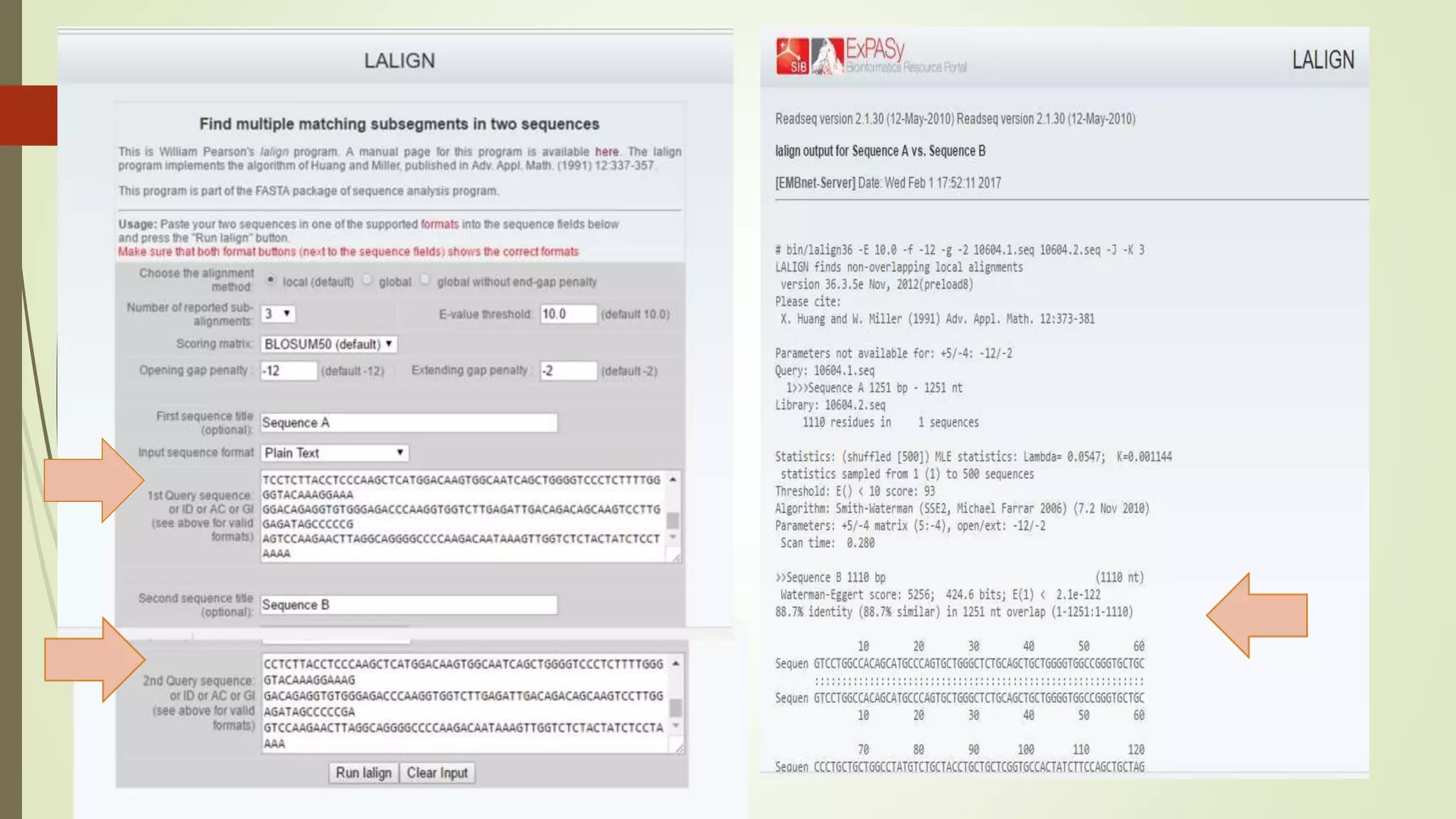
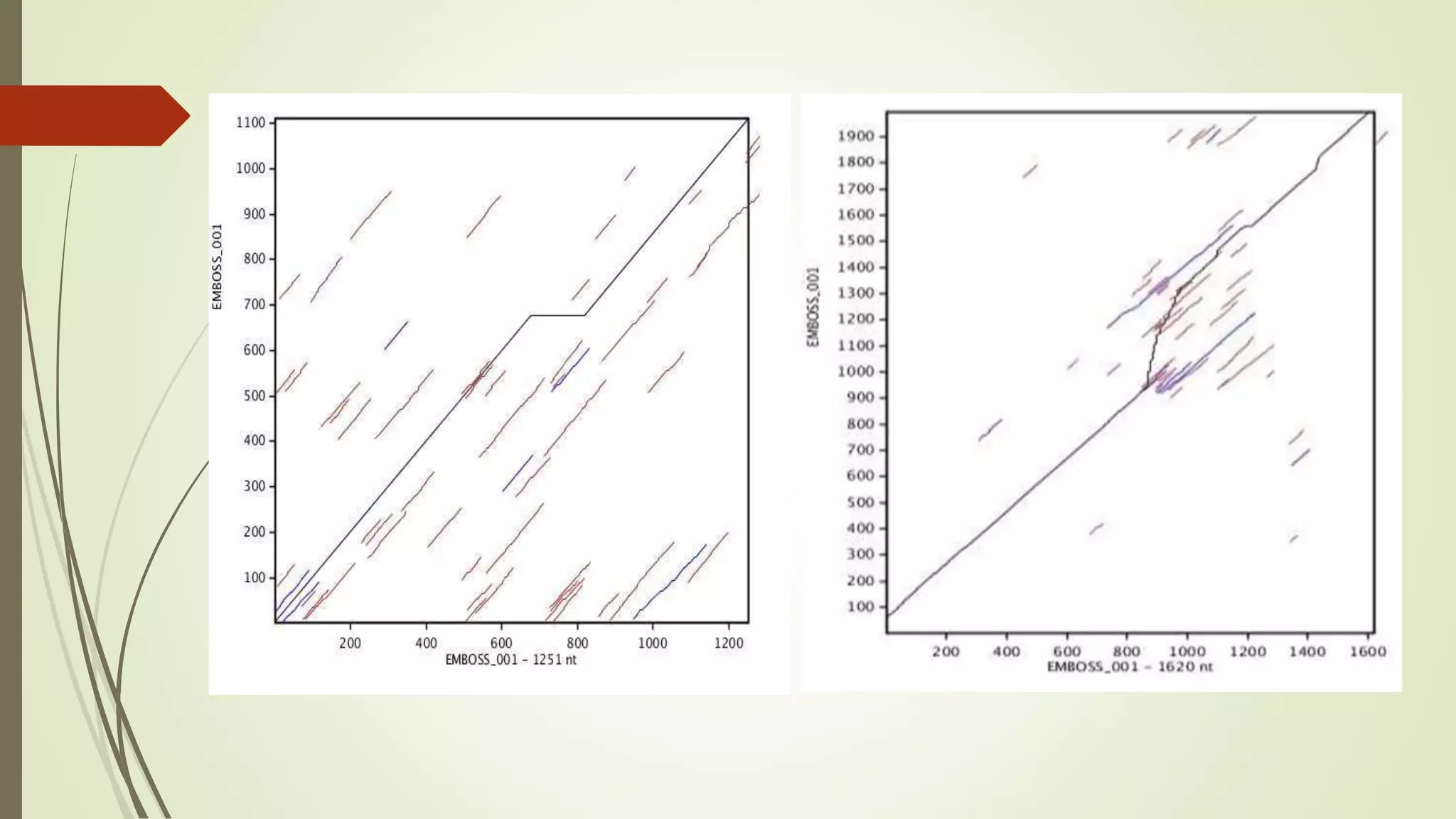
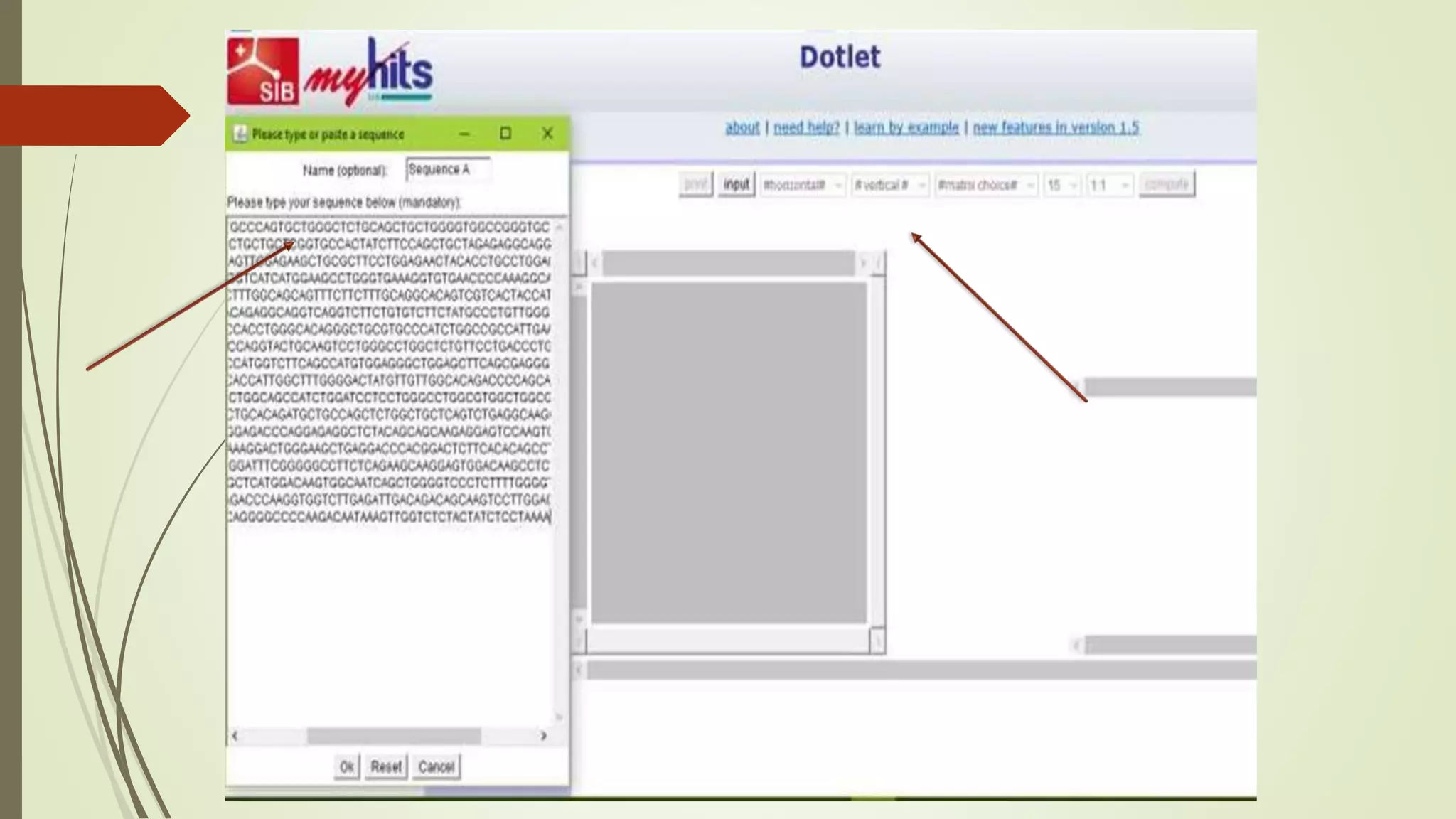
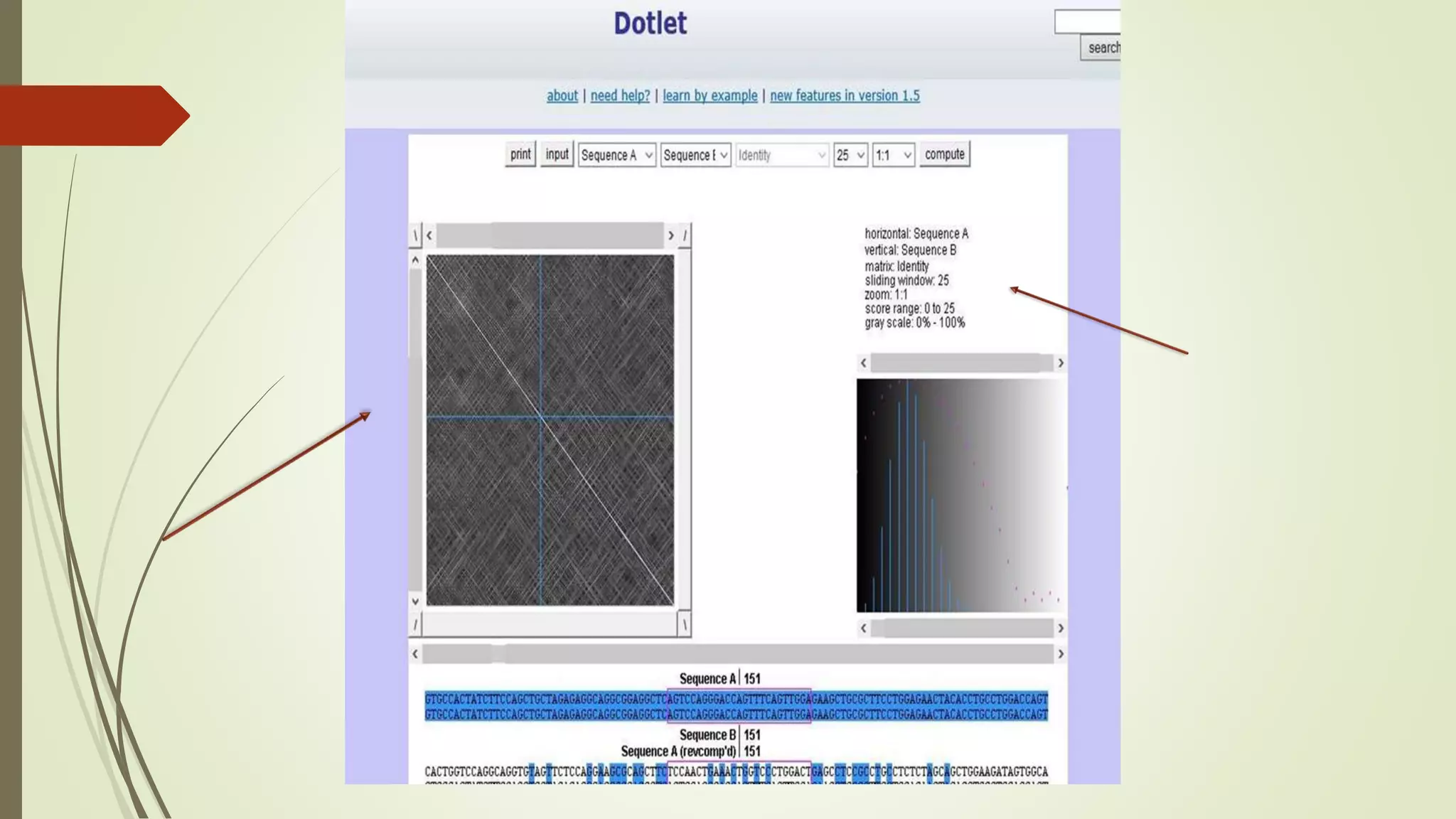
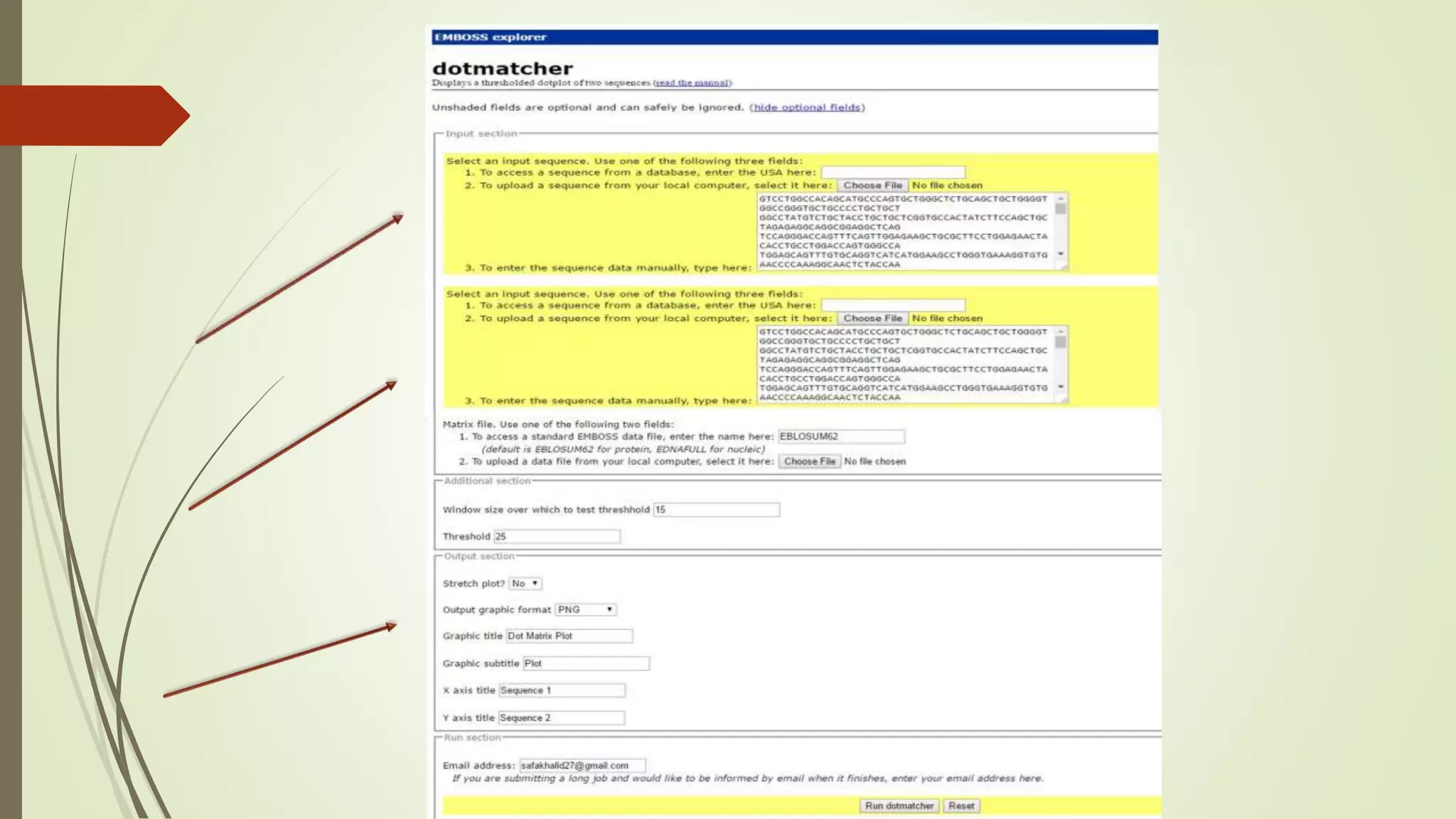
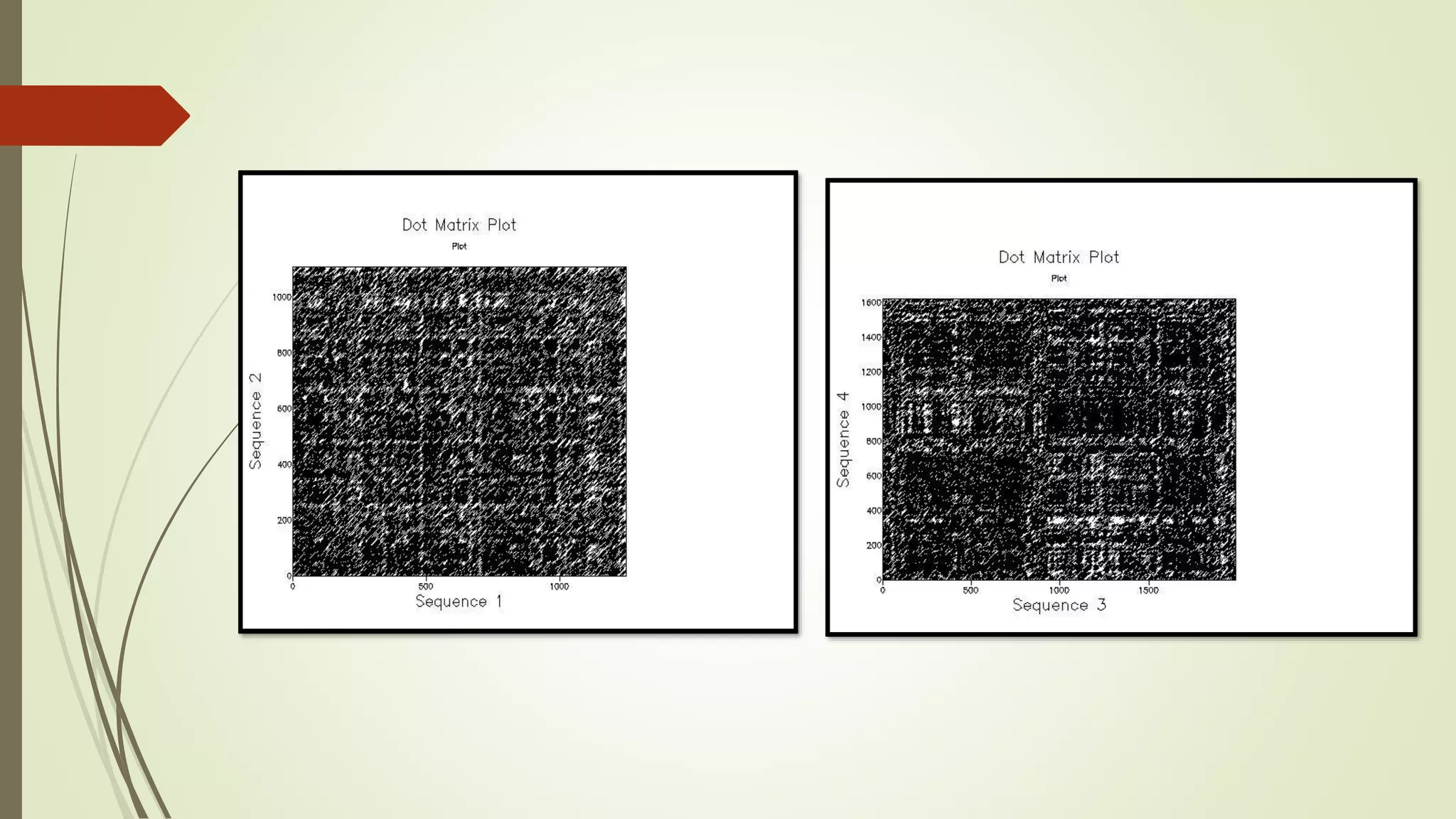
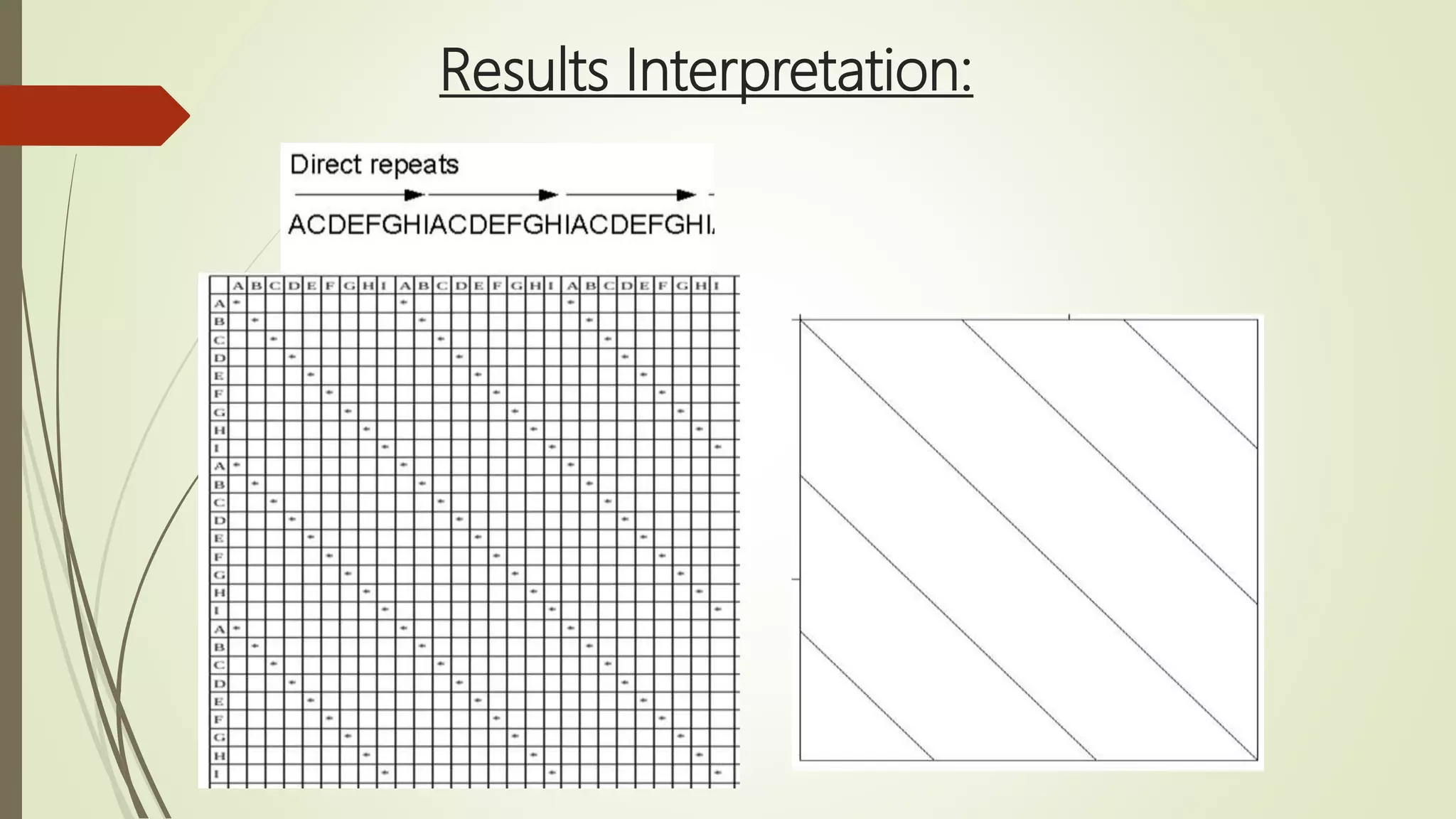
Sequence alignment involves arranging DNA, RNA, or protein sequences to identify similar regions and infer functional or evolutionary relationships. Dot matrix alignment visually represents the similarities between two sequences in a grid, where dots indicate matching characters. Software like LALIGN, DOTLET, and DOTMATCHER can perform dot matrix alignments, calculating scores based on factors like gap penalties to identify significant matches despite differences. Dot plots can reveal similarities like inverted repeats or palindromic sequences.