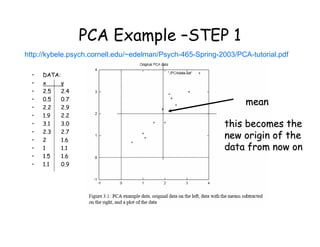
Principal Components Analysis (PCA) is a technique used to simplify complex datasets. It works by transforming the data to a new coordinate system where the greatest variance lies on the first axis (called the first principal component), second greatest variance on the second axis, and so on. This allows for dimensionality reduction by removing later principal components with less variance. PCA is commonly used for applications like face recognition, image compression, and finding patterns in high-dimensional data.


















![PCA Theorem
Let Q = X X
Let Q = X XT
T be the N x N matrix:
be the N x N matrix:
Notes:
Notes:
1.
1. Q is square
Q is square
2.
2. Q is symmetric
Q is symmetric
3.
3. Q is the
Q is the covariance
covariance matrix [aka scatter matrix]
matrix [aka scatter matrix]
4.
4. Q can be very large
Q can be very large (in vision, N is often the number of
(in vision, N is often the number of
pixels in an image!)
pixels in an image!)](https://image.slidesharecdn.com/covariance-221009122304-24b470d5/85/Covariance-pdf-19-320.jpg)