Embed presentation
Downloaded 30 times

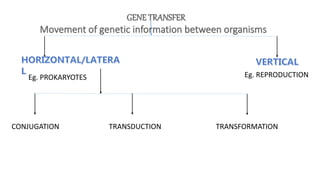
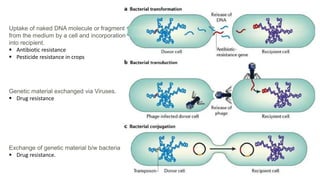
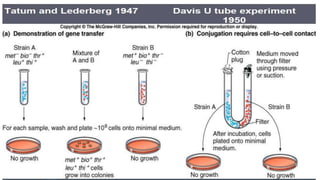

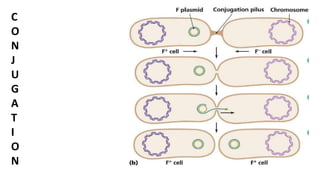
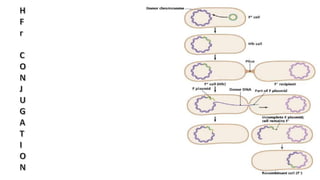

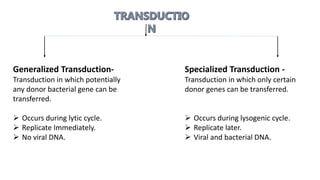

The document discusses molecular genetics in prokaryotes, focusing on mechanisms of genetic material exchange such as conjugation, transformation, and transduction. It explains how these processes can lead to drug and antibiotic resistance in bacteria. Additionally, it outlines the differences between generalized and specialized transduction, including the types of bacteria involved and the conditions of each process.








