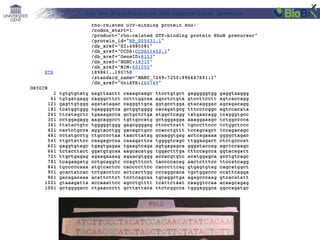
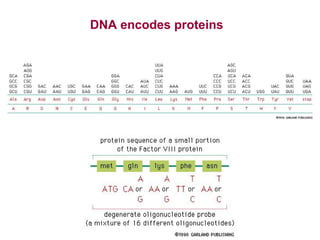
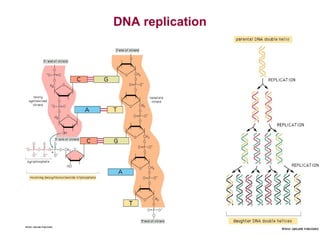
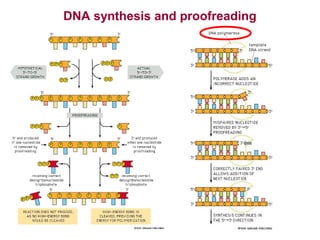
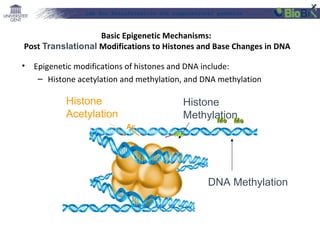
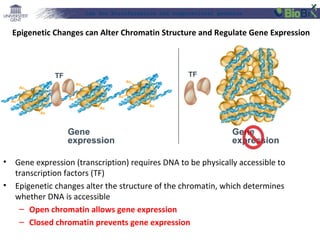
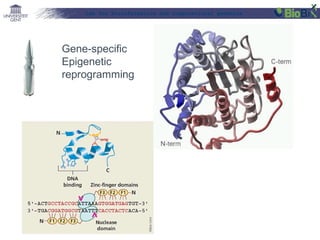
This document provides an overview of bioinformatics and computational genomics. It discusses key topics including DNA structure and function, genetic code, DNA replication, mutations, epigenetics, chromatin structure, histone modifications, DNA methylation, cancer stem cells, personalized medicine using biomarkers, and molecular profiling. The document contains diagrams explaining concepts like DNA packaging into chromatin, basic epigenetic mechanisms involving histone modifications and DNA methylation, and how epigenetic changes can alter chromatin structure and regulate gene expression.

![Lab for Bioinformatics and computational genomics
Overview
^[now][transl⎮comput]ational[epi]genomic$
•
•
•
•
Who ? Where ?
Bioinformatics
(Epi)genetics
Technology: Next Gen
Sequencing
• Personal Genomics](https://image.slidesharecdn.com/20131023dnafordummiesvpresented-131026022933-phpapp01/85/2013-10-23_dna_for_dummies_v_presented-2-320.jpg)







![Lab for Bioinformatics and computational genomics
Overview
^[now][transl⎮comput]ational[epi]genomic$
•
•
•
•
Who ? Where ?
Bioinformatics
(Epi)genetics
Technology: Next Gen
Sequencing
• Personal Genomics](https://image.slidesharecdn.com/20131023dnafordummiesvpresented-131026022933-phpapp01/85/2013-10-23_dna_for_dummies_v_presented-18-320.jpg)















![Lab for Bioinformatics and computational genomics
Overview
^[now][transl⎮comput]ational[epi]genomic$
•
•
•
•
Who ? Where ?
Bioinformatics
(Epi)genetics
Technology: Next Gen
Sequencing
• Personal Genomics](https://image.slidesharecdn.com/20131023dnafordummiesvpresented-131026022933-phpapp01/85/2013-10-23_dna_for_dummies_v_presented-53-320.jpg)
![Lab for Bioinformatics and computational genomics
Overview
^[now][transl⎮comput]ational[epi]genomic$
•
•
•
•
Who ? Where ?
Bioinformatics
(Epi)genetics
Technology: Next Gen
Sequencing
• Personal Genomics](https://image.slidesharecdn.com/20131023dnafordummiesvpresented-131026022933-phpapp01/85/2013-10-23_dna_for_dummies_v_presented-55-320.jpg)