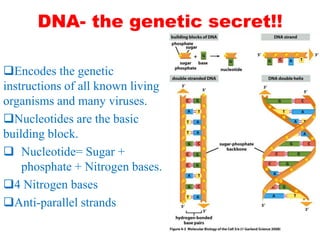
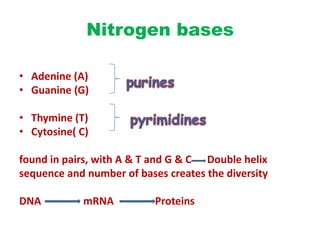
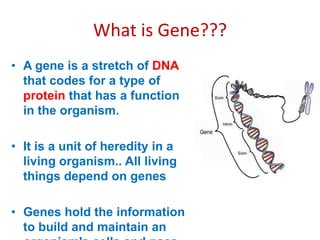
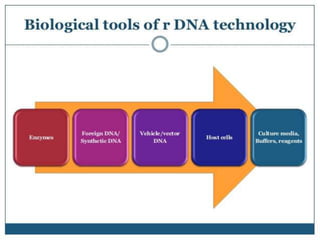
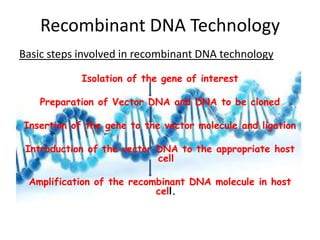
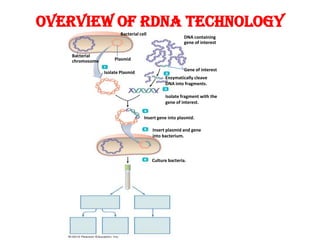
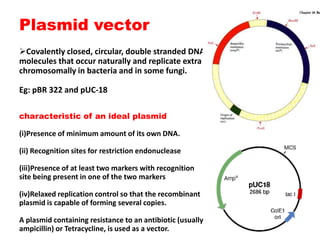
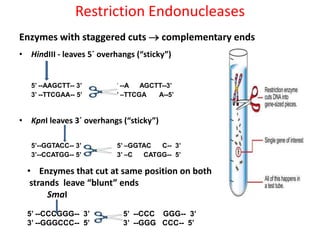
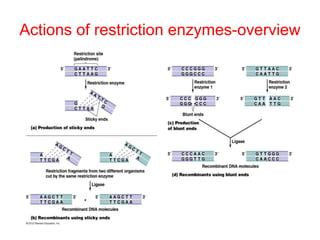
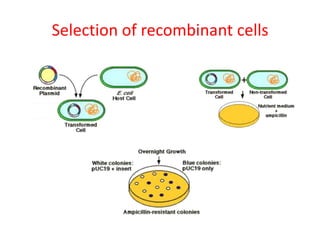
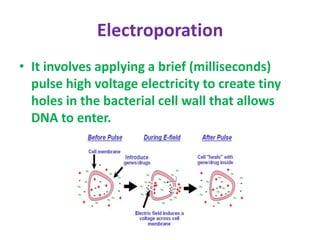
This document provides an overview of recombinant DNA technology. It begins by describing the basic components and structure of DNA, including nucleotides, nitrogen bases, and how DNA encodes genetic instructions. It then defines what a gene is and explains that recombinant DNA technology involves joining DNA fragments from different sources. The key steps are described as isolating the gene of interest, inserting it into a vector like a plasmid, introducing the vector into a host cell, and amplifying the recombinant DNA. A variety of applications are mentioned, such as producing pharmaceuticals, genetically modifying crops, and using bacteria to break down environmental waste.