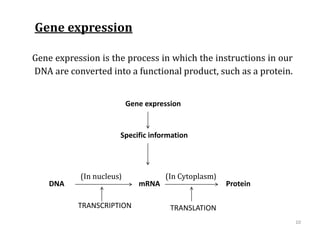
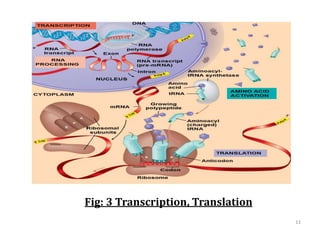
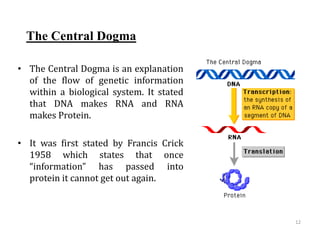
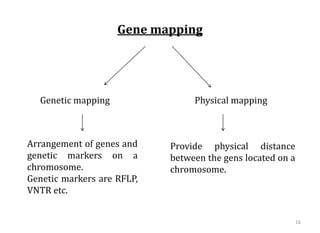
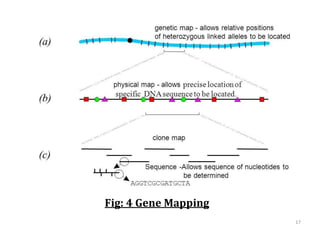
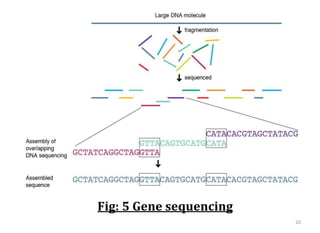
Genome organization refers to the sequential arrangement of genes within an organism. The genome consists of DNA packaged into chromosomes, which contain genes that code for proteins and RNA. Gene expression involves transcription of DNA into mRNA and translation of mRNA into proteins. The human genome project mapped the entire human genome sequence to further understand gene function and human health. Genome sequencing and mapping are important for disease diagnosis, drug development, and other medical applications.