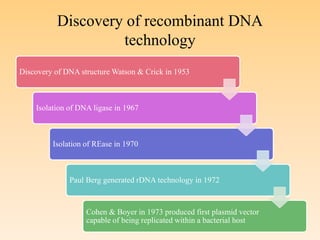
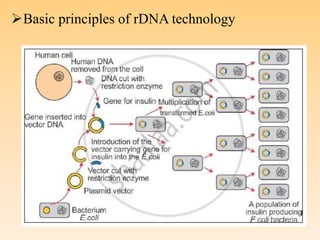
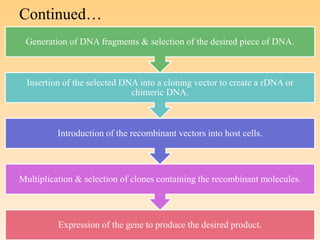
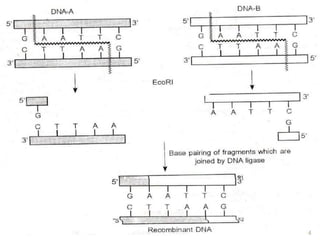
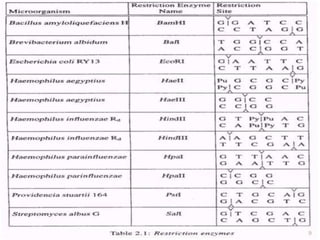
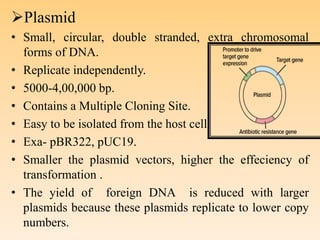
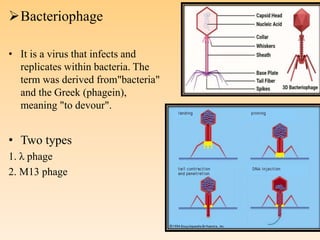
Recombinant DNA technology allows DNA from different species to be isolated, cut, spliced together, and multiplied. This produces recombinant molecules with DNA from different sources. Key steps include generating DNA fragments, inserting a fragment into a cloning vector, and introducing the recombinant vector into a host cell for expression. Restriction enzymes are important tools that cut DNA at specific recognition sequences. Common vectors used include plasmids, bacteriophages, and artificial chromosomes in bacteria or yeast. DNA ligase joins DNA strands together, facilitating recombination. Applications include pharmaceuticals, disease diagnosis, forensics, agriculture, and more.