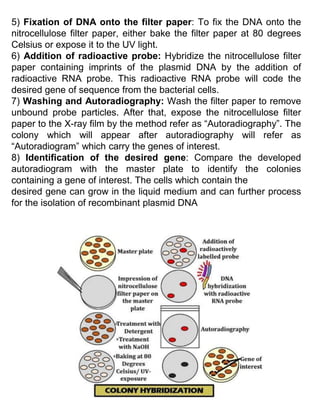
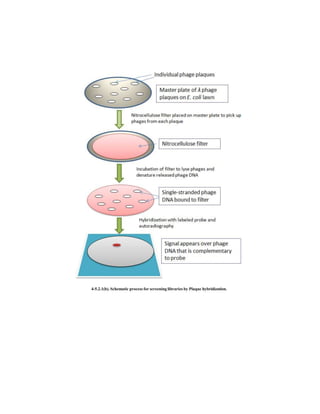
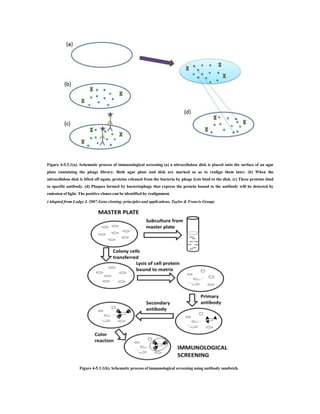
The document provides an overview of screening and preservation methods for DNA libraries, focusing on techniques for identifying clones containing a gene of interest. Key methods include hybridization (colony and plaque hybridization) and immunological screening, each with its own procedural steps and advantages. Additionally, it discusses functional complementation for the isolation of missing functions in mutant cells and highlights the applications and drawbacks of these screening methods.