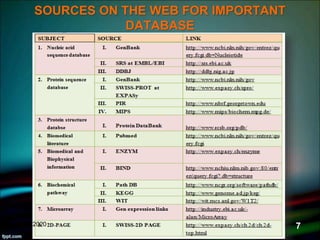
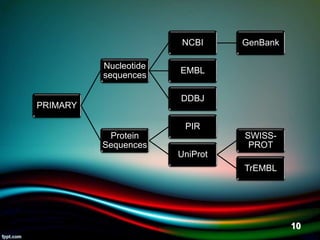
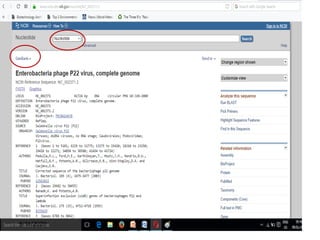
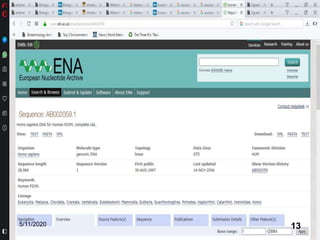
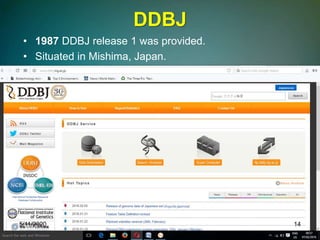
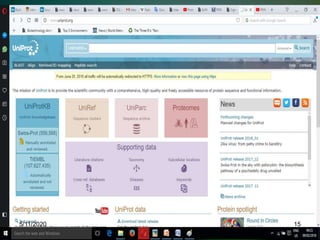
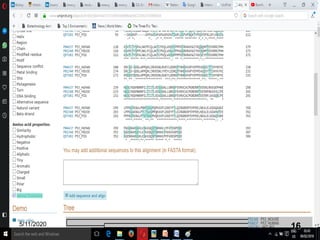
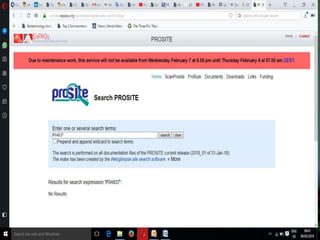
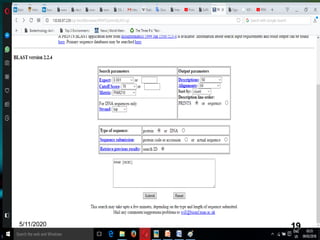
The document discusses biological databases, explaining their definitions, types (primary, secondary, composite, and tertiary), and the importance of bioinformatics in managing biological data. It highlights various primary nucleotide and protein sequence databases such as NCBI, GenBank, and EMBL-EBI, while also covering secondary databases including Prosite and Prints. The conclusion emphasizes the necessity of storing and analyzing extensive biological data to aid scientific research.