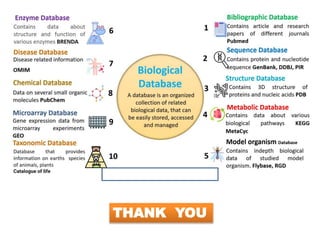
The document discusses biological databases in bioinformatics, explaining their purpose, structure, and types including primary, secondary, and composite databases. Primary databases store original biological data while secondary databases provide processed information derived from primary data. The major public sequence databases mentioned are GenBank, EMBL, and DDBJ, which house raw nucleic acid sequence data from researchers worldwide.