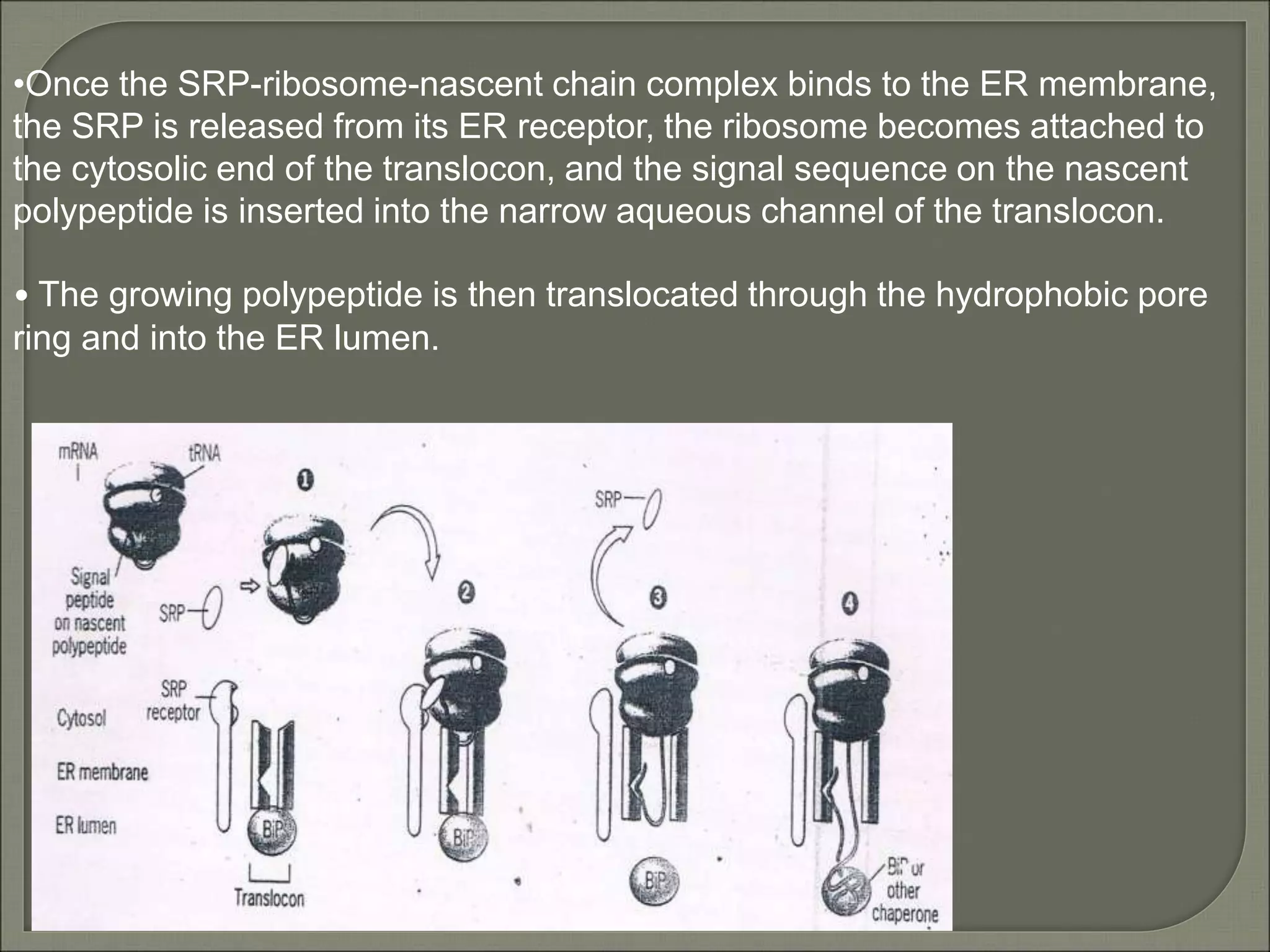
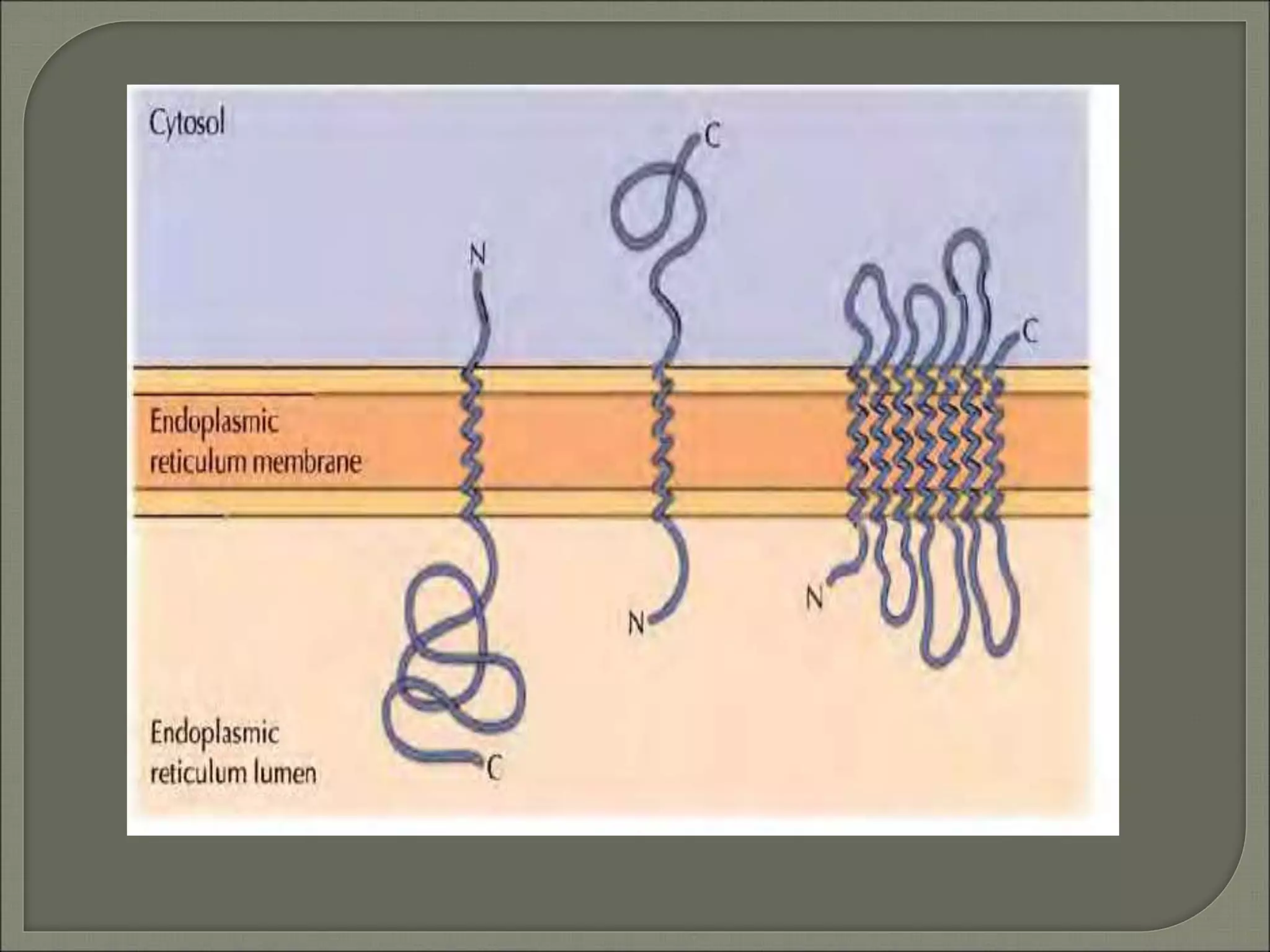
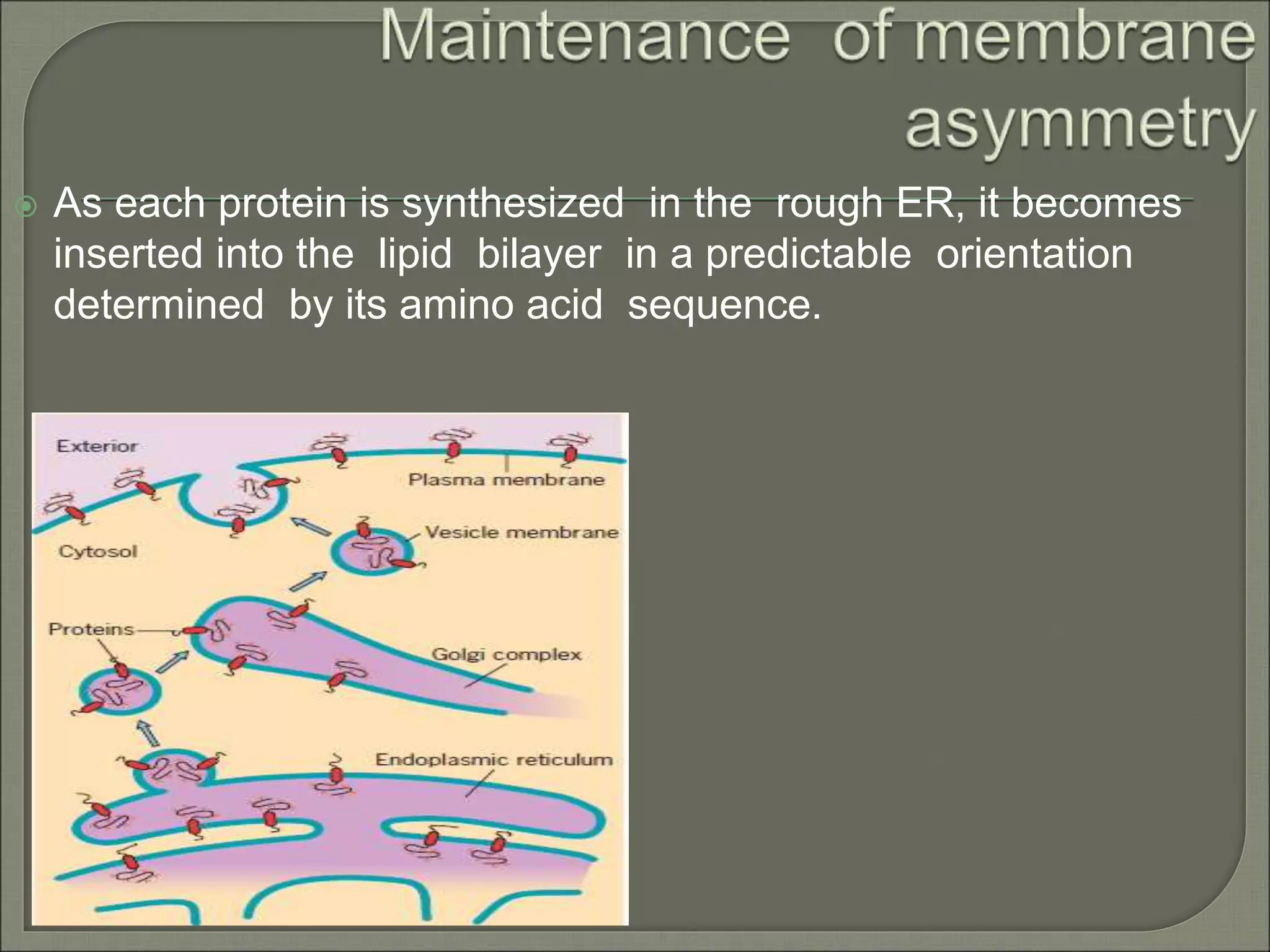
The document discusses the processes of protein synthesis, particularly focusing on the synthesis of secretory and integral membrane proteins in the rough endoplasmic reticulum (ER). Key processes such as transcription, translation, and the roles of signal recognition particles and molecular chaperones in protein processing are highlighted. The synthesis of membrane proteins, their integration into the lipid bilayer, and their functional roles in the cell are also covered.