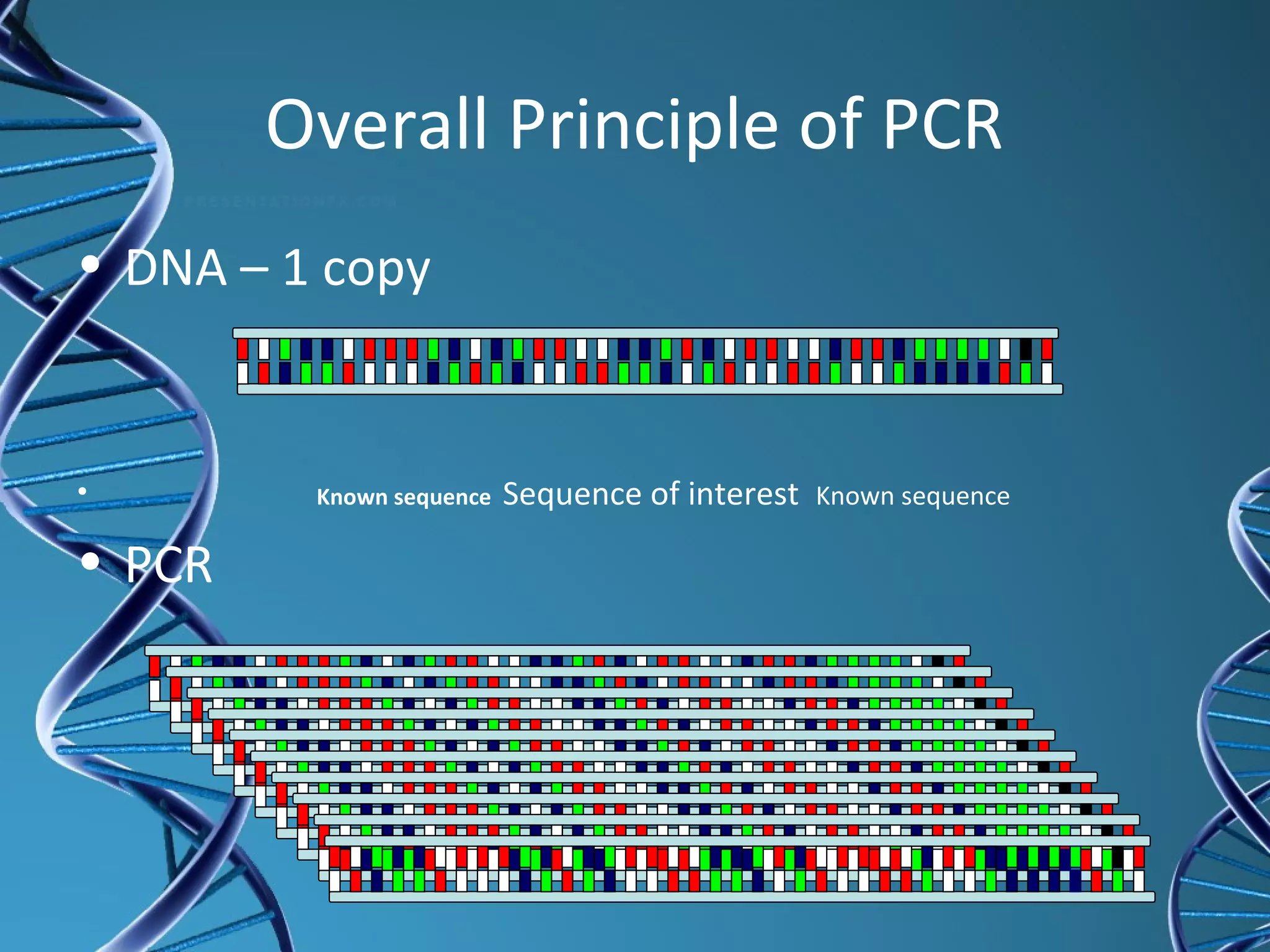
The document provides an overview of the polymerase chain reaction (PCR), detailing its history, principles, components, and applications. Developed by Dr. Kary Mullis in the 1980s, PCR amplifies specific DNA sequences for various scientific and medical applications, including genetic testing and forensic analysis. Key advantages include the requirement of small DNA amounts, faster results, and high precision in analyzing genetic material.