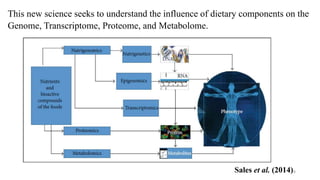
This document provides an overview of nutrigenomics, including:
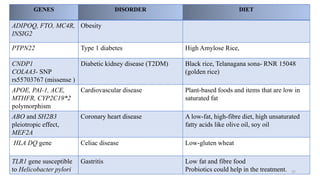
- The definition of nutrigenomics as studying the relationship between human genome, nutrition and health.
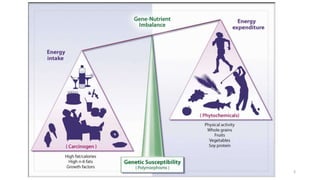
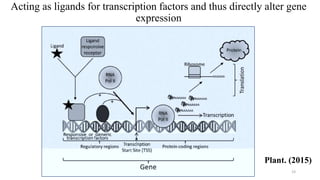
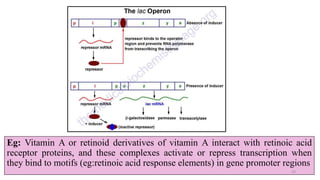
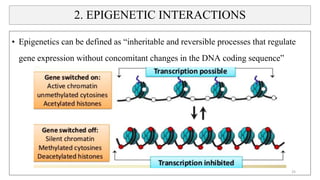
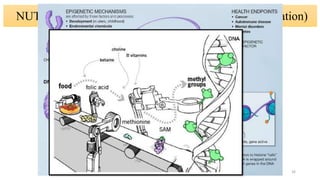
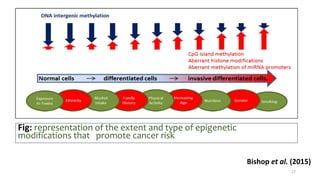
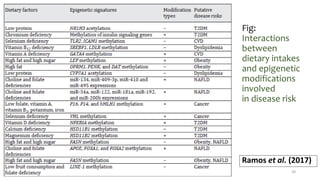
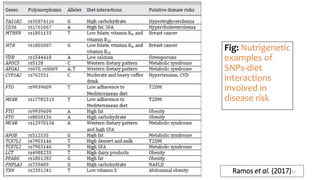
- How nutrients can interact with genes through direct interaction, epigenetic interactions, and genetic variations.
- Tools and databases used in nutrigenomics research like BioConductor and the Nutritional Phenotype database.
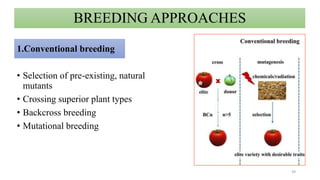
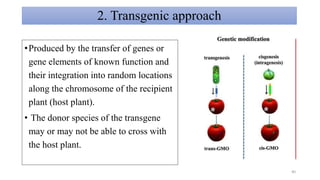
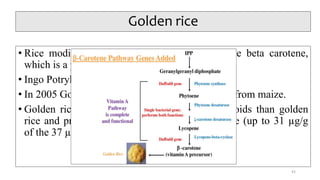
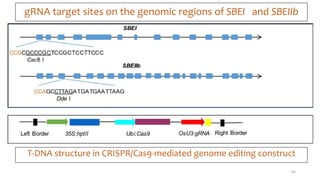
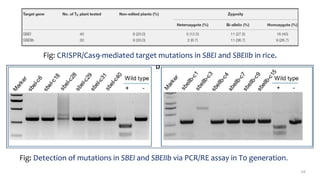
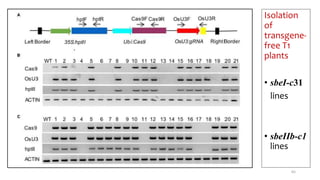
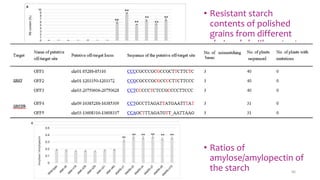
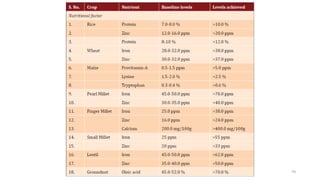
- Examples of how nutrigenomics research can inform plant breeding to develop crops with improved nutritional profiles through approaches like marker-assisted selection and genome editing.