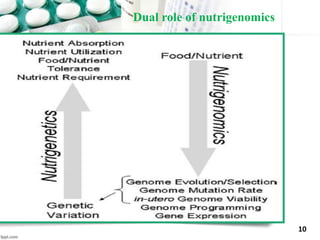
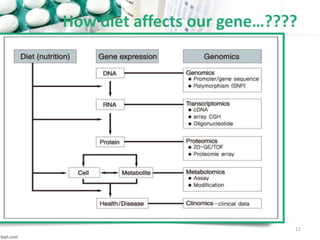
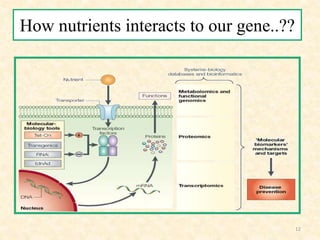
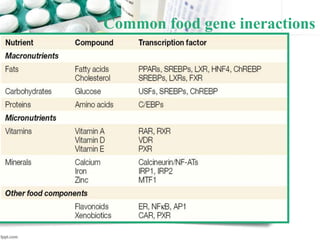
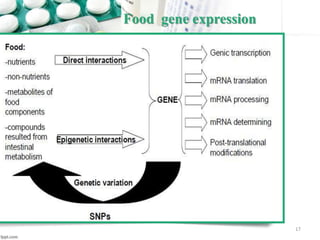
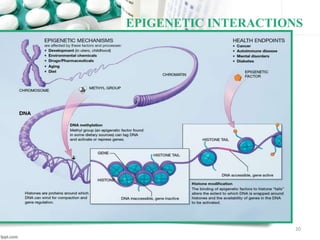
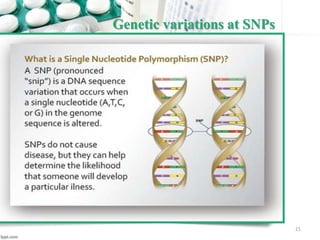
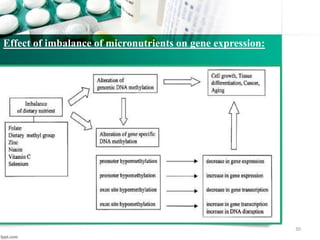
This document provides an introduction to nutrigenomics and its applications. It discusses how nutrigenomics is the study of how dietary components interact with genes and alter gene expression. There are different types of food-gene interactions, including direct interactions where nutrients directly bind to receptors and regulate genes, and epigenetic interactions where nutrients can alter DNA structure and chronically change gene expression. Nutrigenomics helps understand how an individual's genetic makeup can influence their susceptibility to diet-related diseases and how personalized diets based on genetics can be used for disease prevention and treatment. The document outlines several examples of how nutrigenomics provides insights into cardiovascular, cancer, obesity and hypertension by studying genetic factors and their interaction with diet.