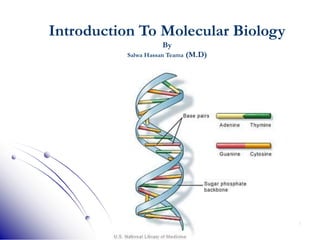
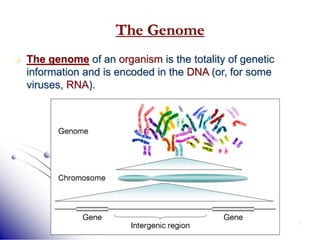
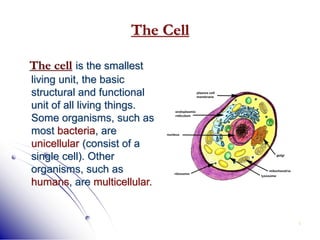
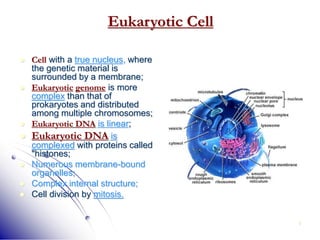
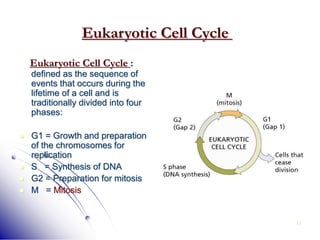
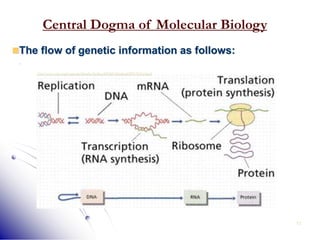
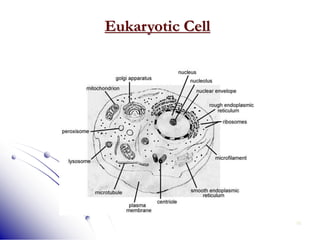
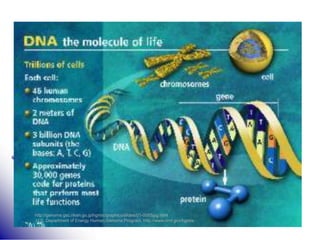
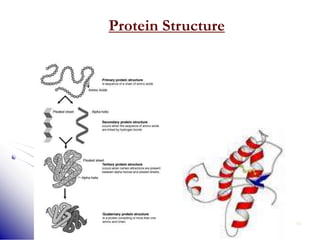
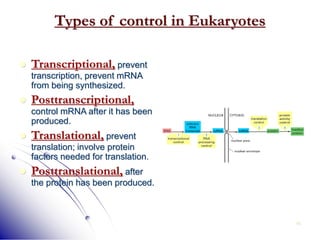
This document provides an introduction to molecular biology. It discusses the key components of cells, including DNA, RNA, chromosomes, and organelles. The central dogma of molecular biology is explained as the flow of genetic information from DNA to RNA to protein. The structures and functions of both eukaryotic and prokaryotic cells are described. DNA replication, transcription, and protein synthesis are summarized as the basic processes by which genetic information is passed from parents to offspring.