This document summarizes a study investigating the enzymatic degradation of lignin model compounds by versatile peroxidase (VP). The key findings are:
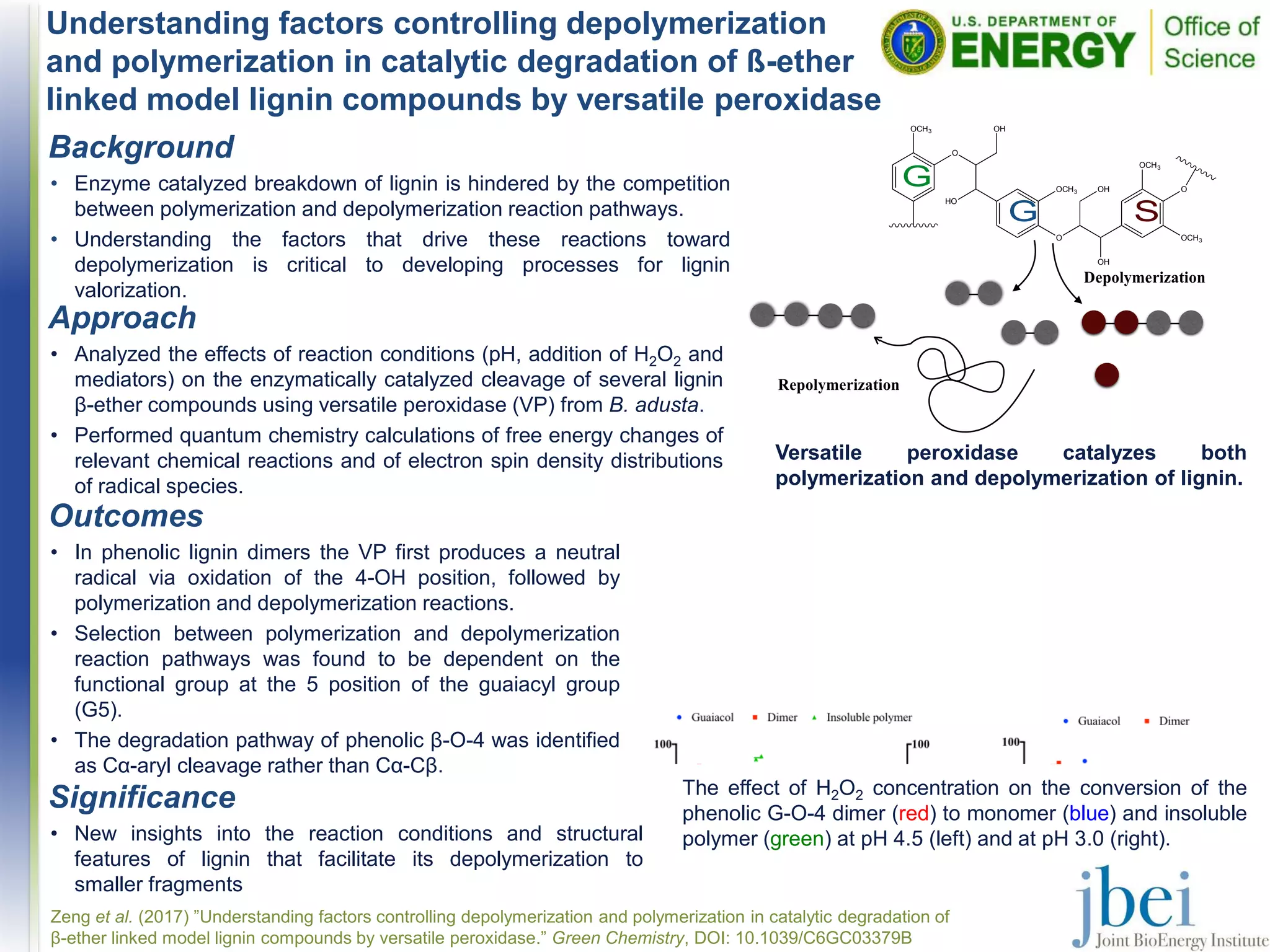
1) VP can catalyze both the polymerization and depolymerization of lignin, with the reaction pathway depending on factors like pH, H2O2 concentration, and the functional groups on the lignin compounds.
2) Degradation of phenolic β-O-4 lignin dimers by VP proceeds through oxidation, followed by competing polymerization or depolymerization reactions depending on conditions.
3) The functional group at the 5 position of guaiacyl units influences whether polymerization or depolymerization occurs.