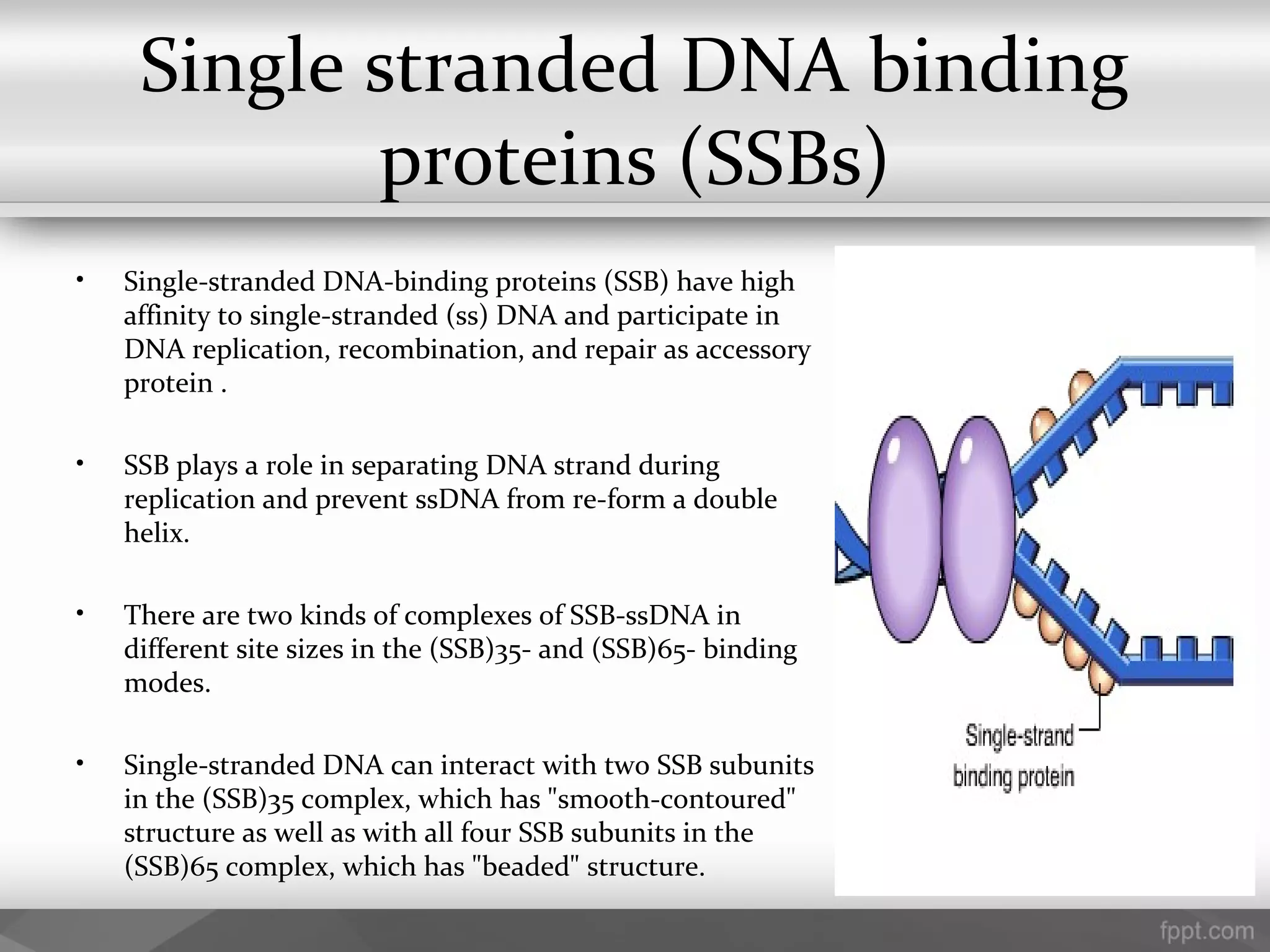
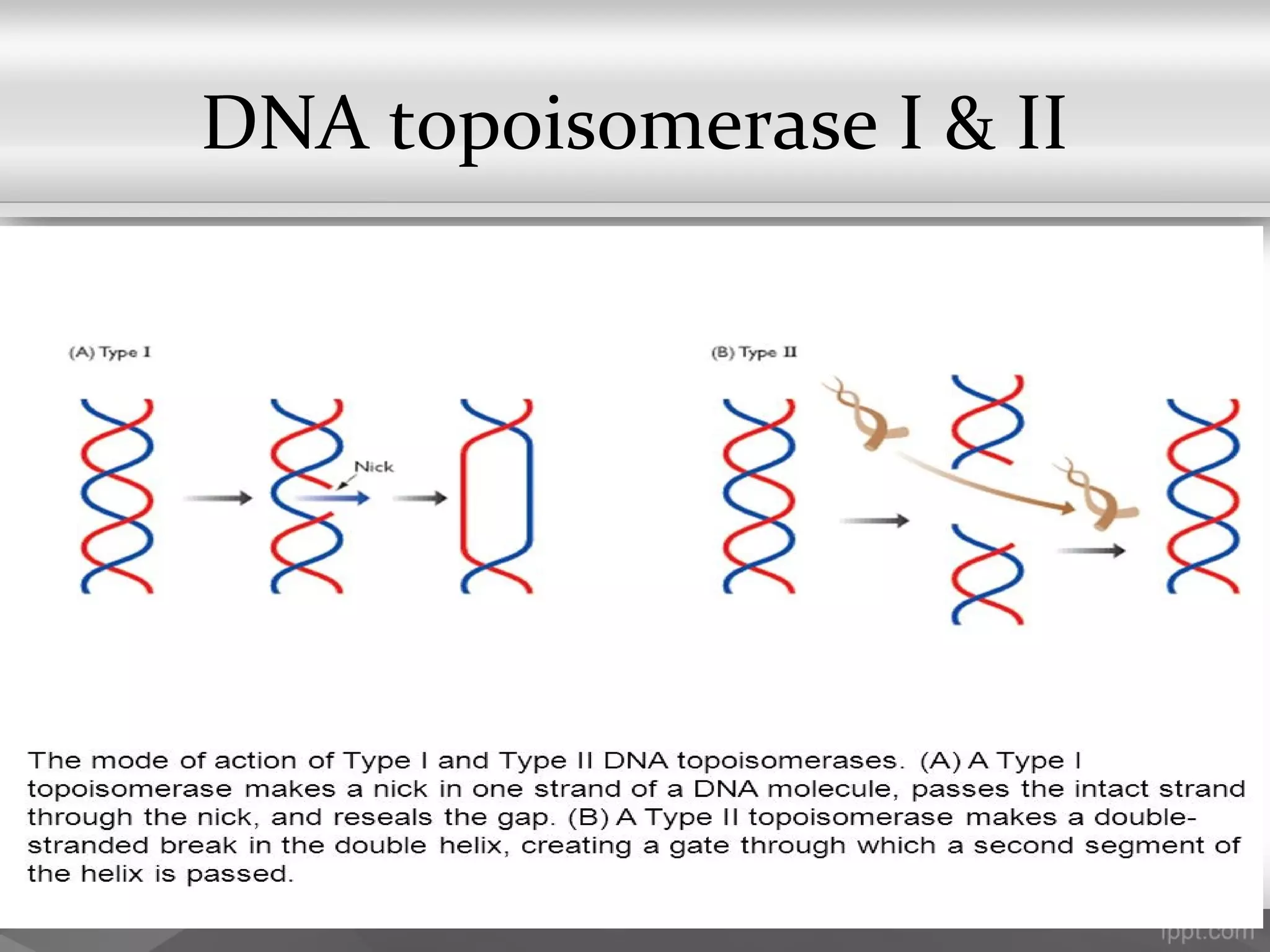
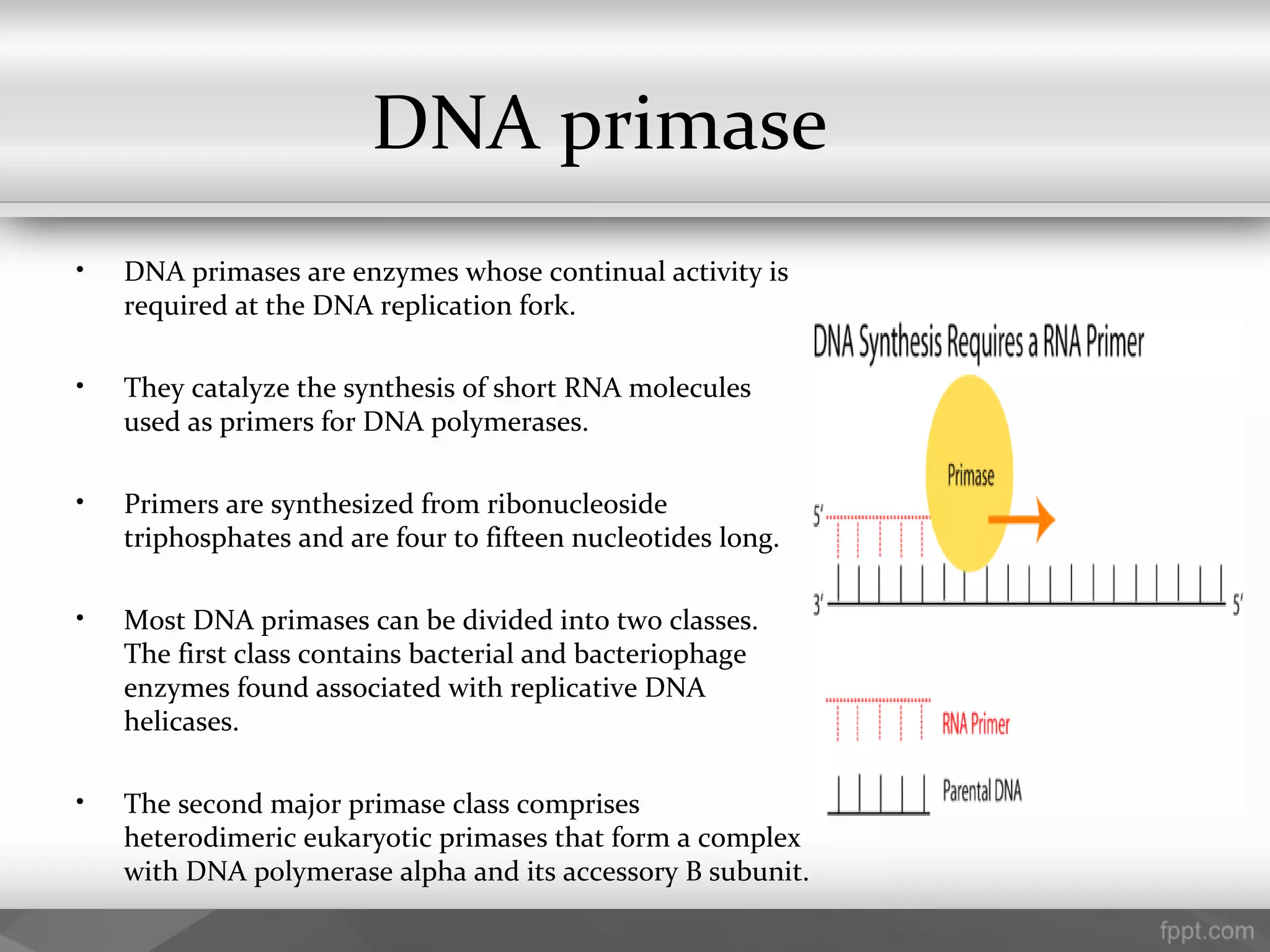
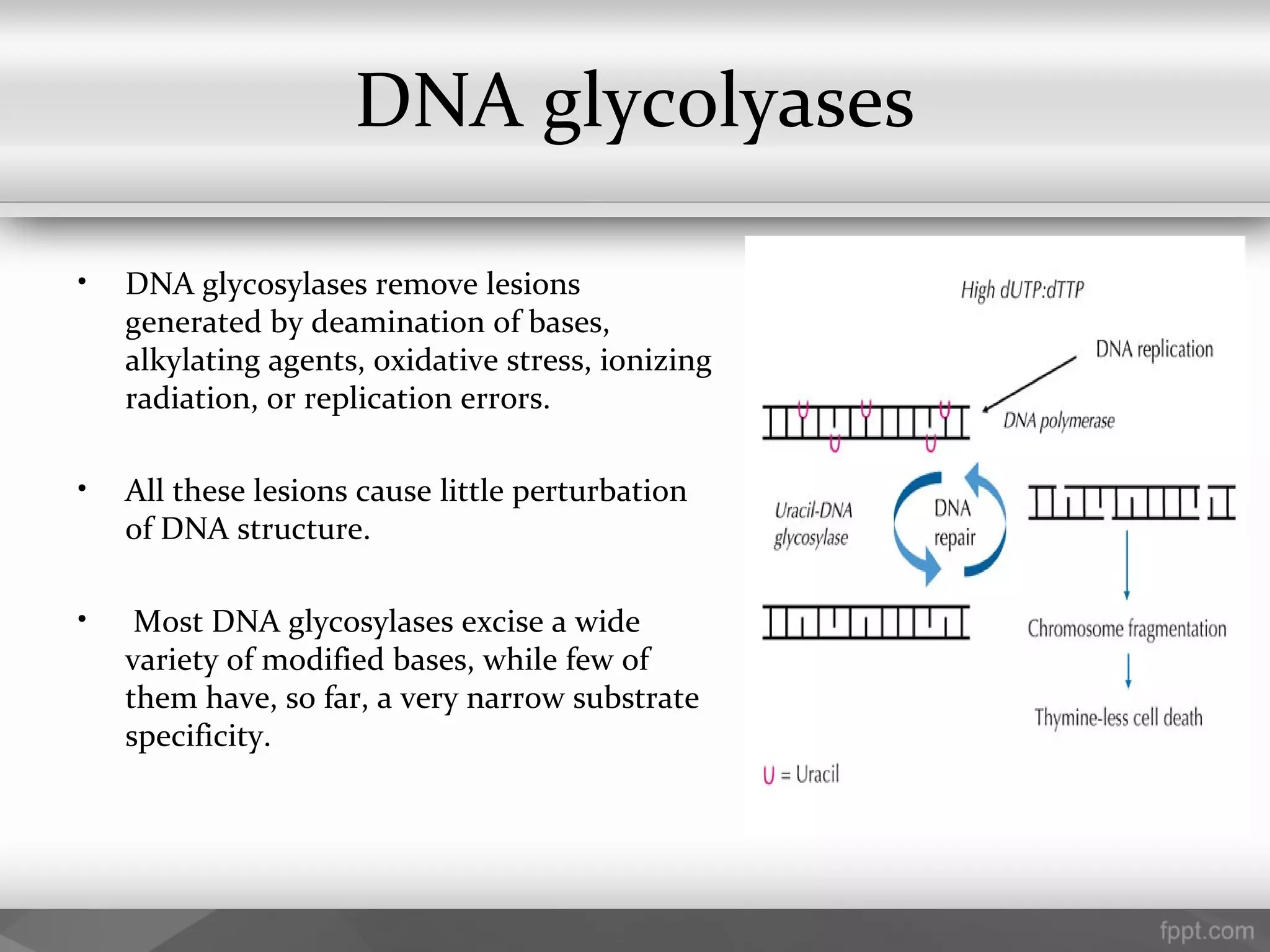
This document summarizes several important components of DNA replication machinery, including DNA helicases, single-stranded DNA binding proteins, DNA topoisomerase, DNA primase, DNA polymerase, DNA ligase, DNA glycolyses, and telomeres. It describes what each component is, its role in DNA replication, and key properties. For example, it notes that DNA helicases use ATP to break hydrogen bonds between nucleotide base pairs, single-stranded DNA binding proteins prevent reformation of the DNA double helix, and DNA ligase reseals nicked DNA strands using ATP.