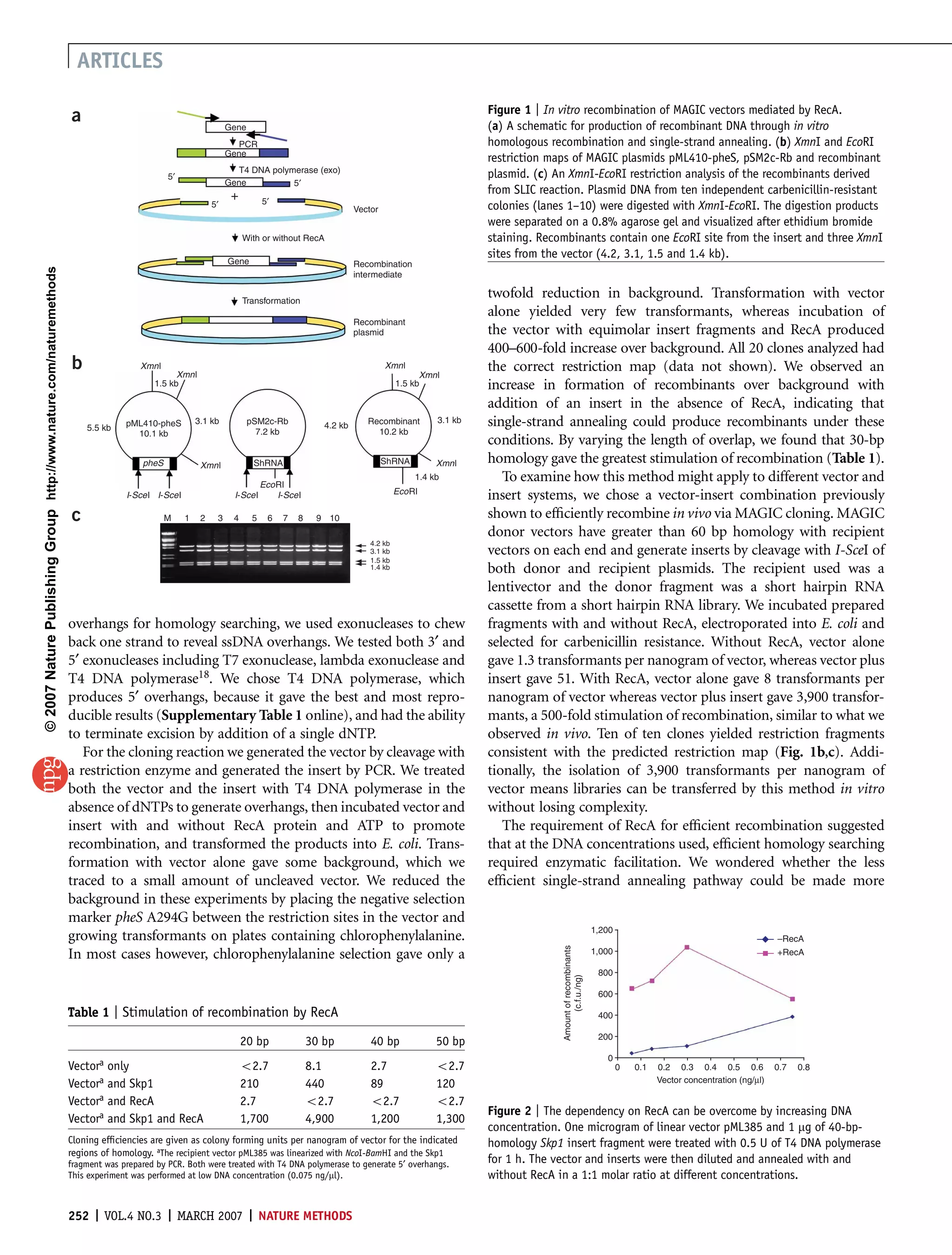
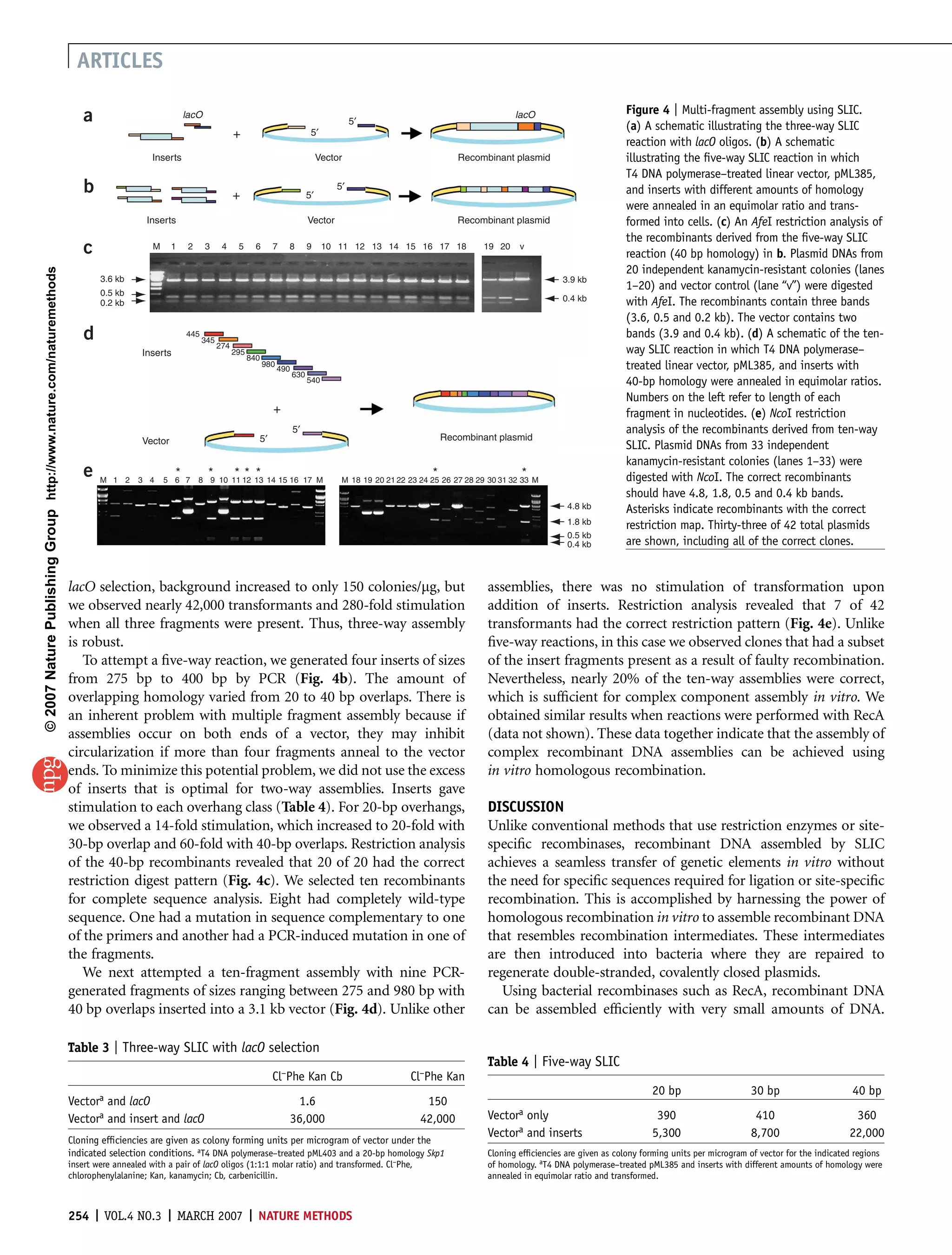
1) The document describes a new cloning method called Sequence and Ligation-Independent Cloning (SLIC) that uses in vitro homologous recombination to assemble multiple DNA fragments into a plasmid.
2) SLIC relies on exonuclease-generated single-stranded DNA overhangs on the vector and insert fragments to facilitate homologous recombination without requiring specific sequences.
3) Experiments show SLIC can efficiently assemble up to 5 or 10 DNA fragments simultaneously and circumvents limitations of other cloning methods by not requiring specific sequences and allowing various insert-vector combinations.