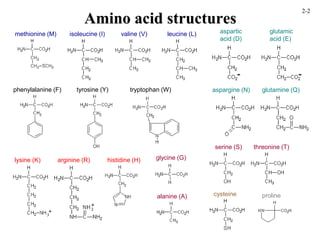
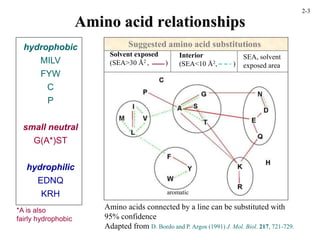
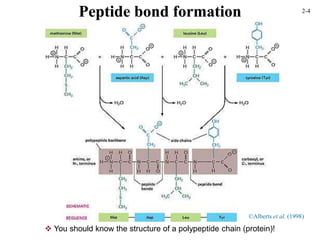
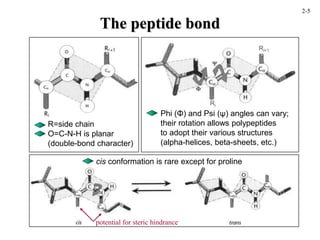
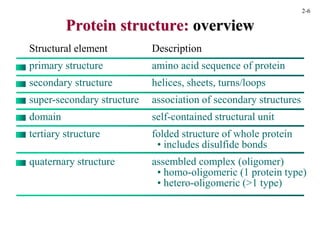
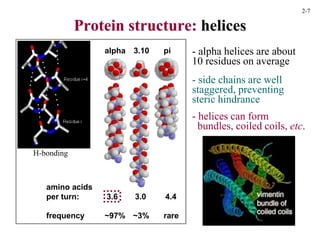
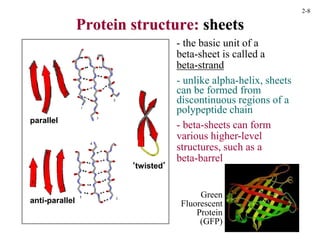
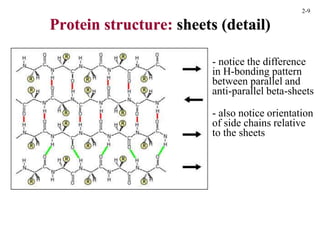
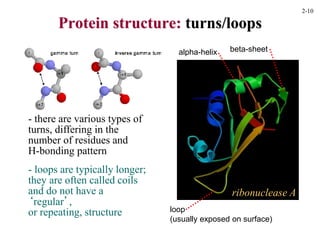
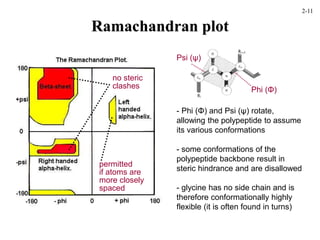
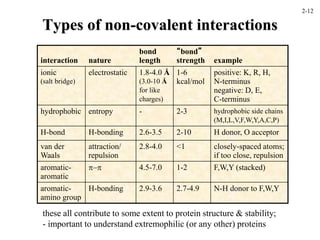
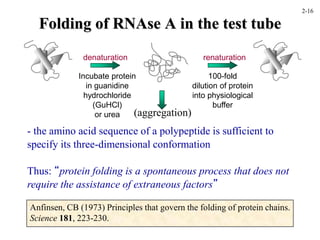
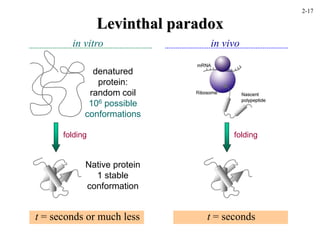
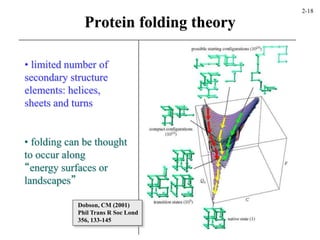
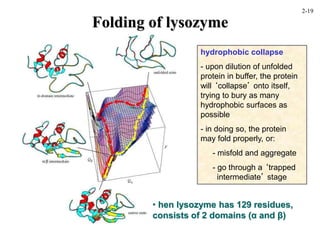
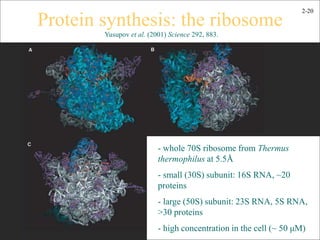
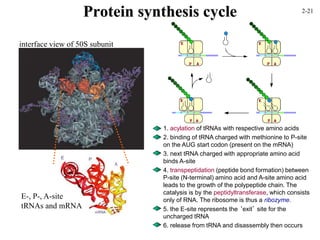
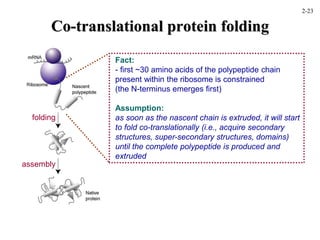
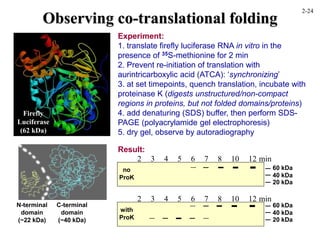
This document discusses protein structure and synthesis. It begins by describing the primary, secondary, tertiary, and quaternary structures of proteins. This includes the structures of alpha helices, beta sheets, turns, and domains. It then discusses protein translation, noting that proteins begin folding as they emerge from the ribosome in a co-translational manner. The final section discusses protein folding and some of the challenges of the folding process.