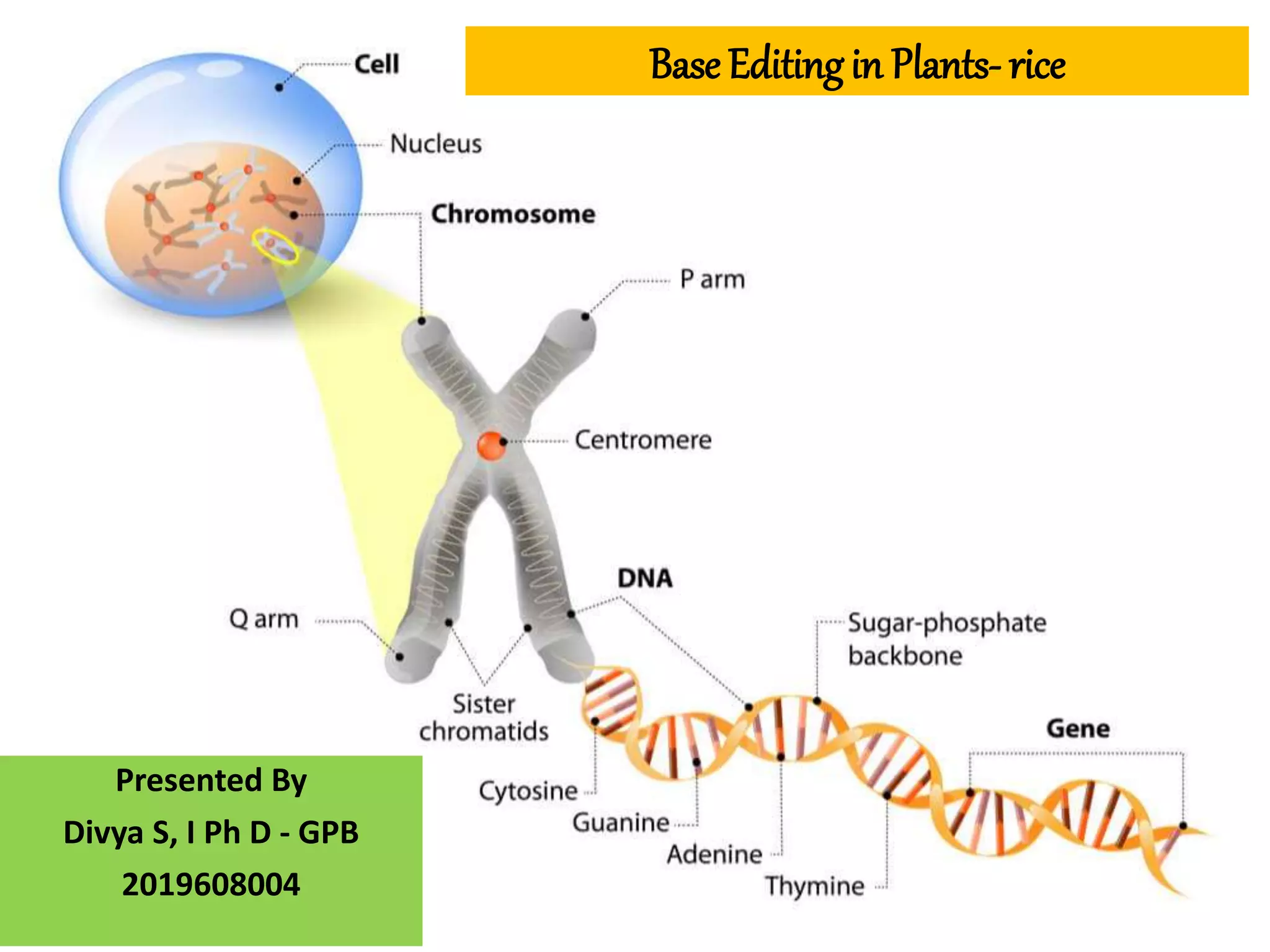
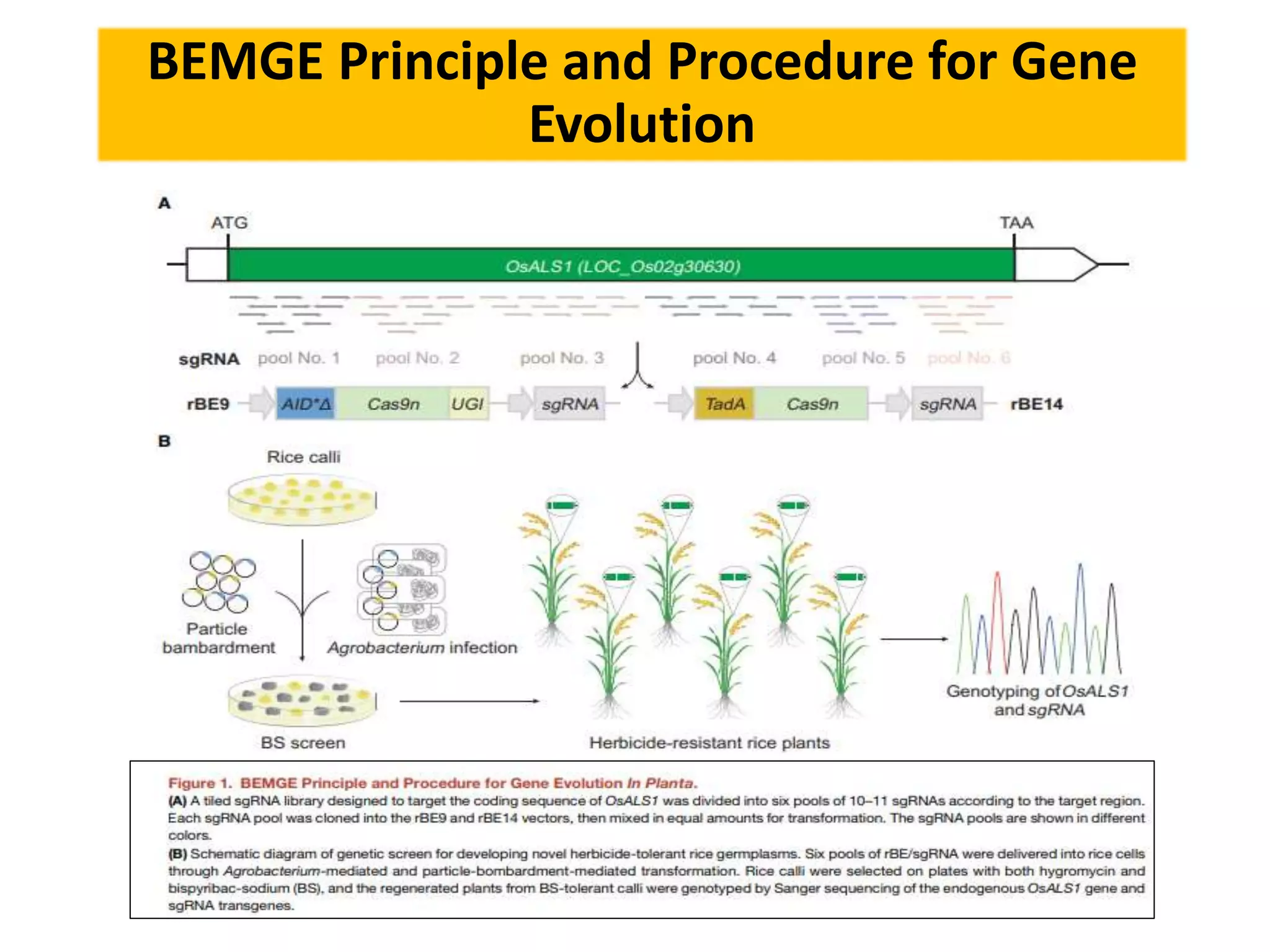
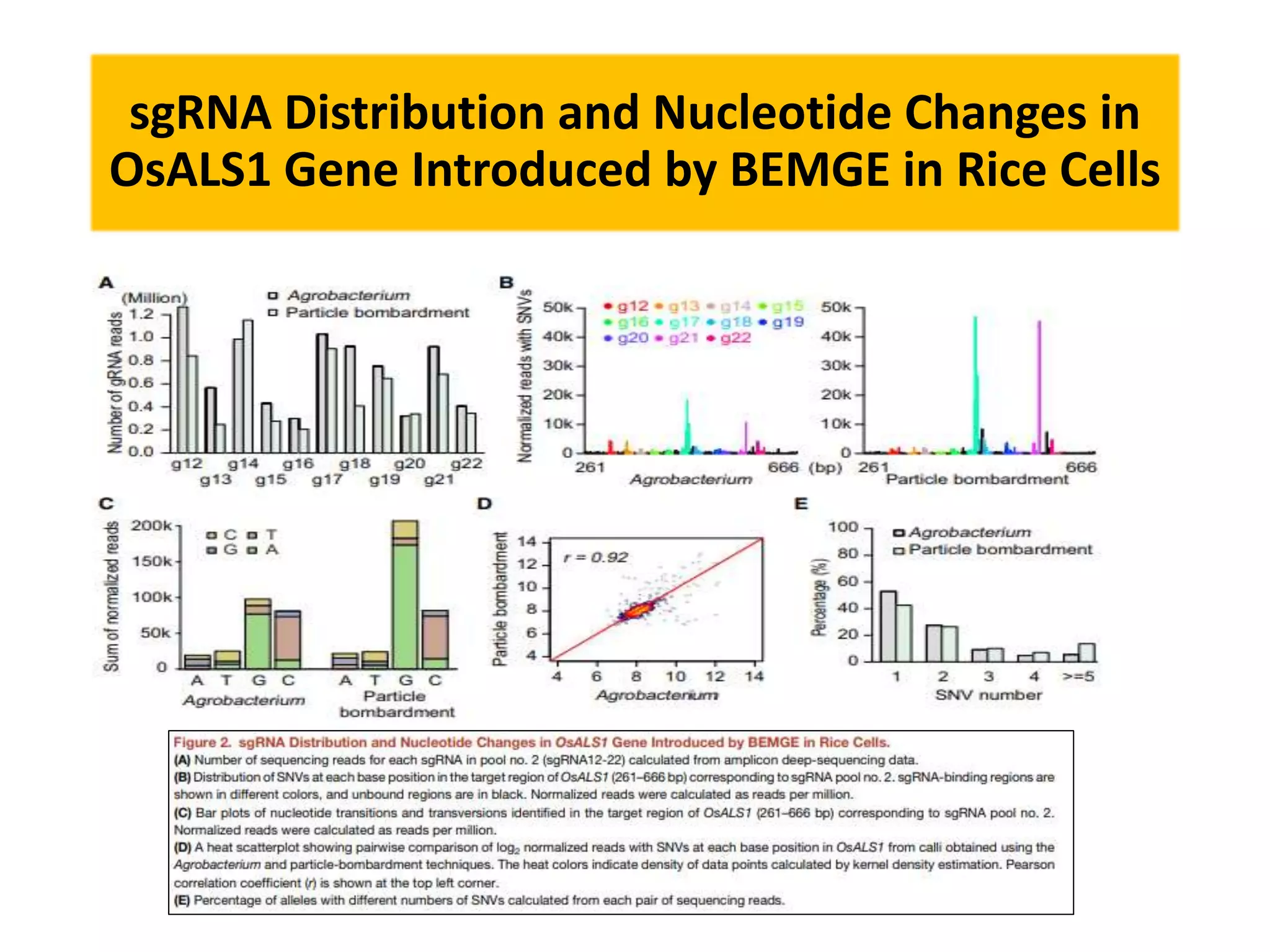
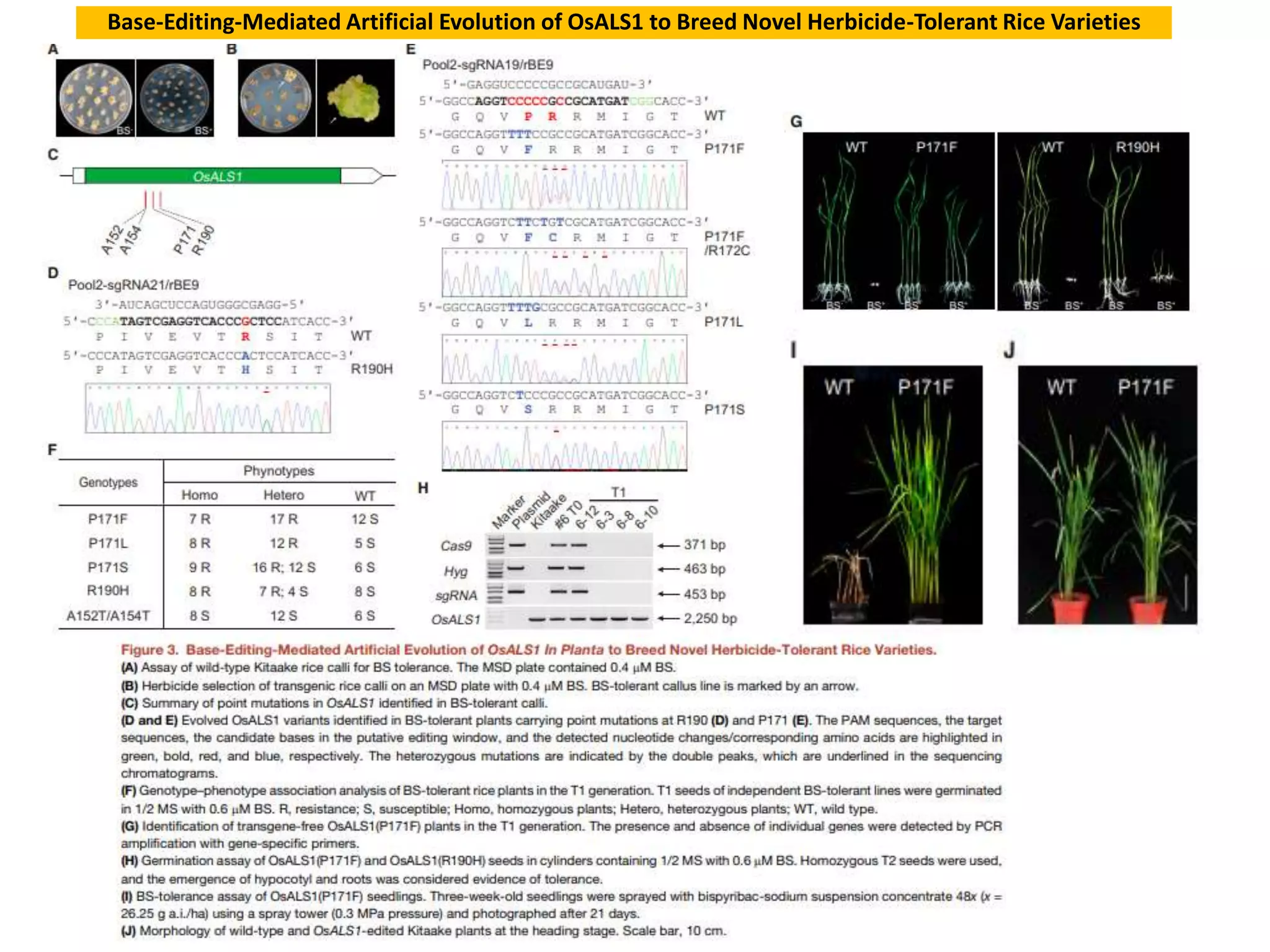
Base editing is being used to induce precise mutations in rice in order to accelerate crop improvement. Researchers developed a base editing-mediated gene evolution method to diversify the sequence of the rice acetolactate synthase 1 (OsALS1) gene, which codes for the target of herbicide resistance. Multiple sgRNAs were used to introduce mutations across the OsALS1 locus. Novel mutations were identified that conferred resistance to the herbicide bispyribac-sodium. The resistant mutation was then introduced into an elite rice cultivar through base editing to generate a new herbicide tolerant variety. This demonstrates how base editing can be used to artificially evolve genes and introduce beneficial traits into commercial crops.