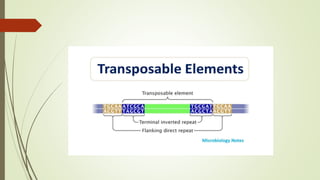
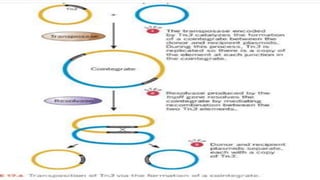
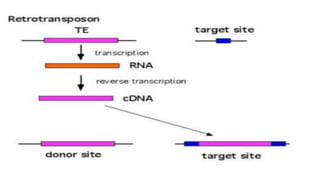
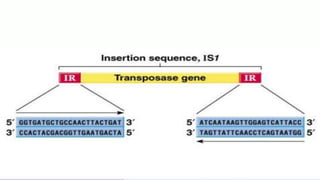
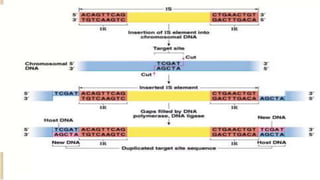
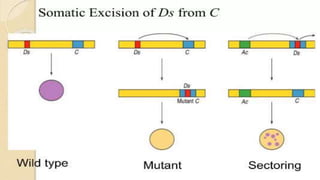
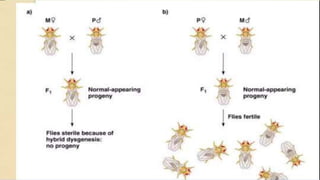
Transposable elements are DNA sequences that can change positions within a genome. They were first discovered by Barbara McClintock in corn in the 1940s. There are three main mechanisms of transposition: replicative, conservative, and retrotransposition. Transposable elements are found in both prokaryotes and eukaryotes and can cause mutations, gene alterations, and genetic diversity. They have applications in mutagenesis and gene expression studies but can also contribute to genetic diseases.