Embed presentation
Download to read offline


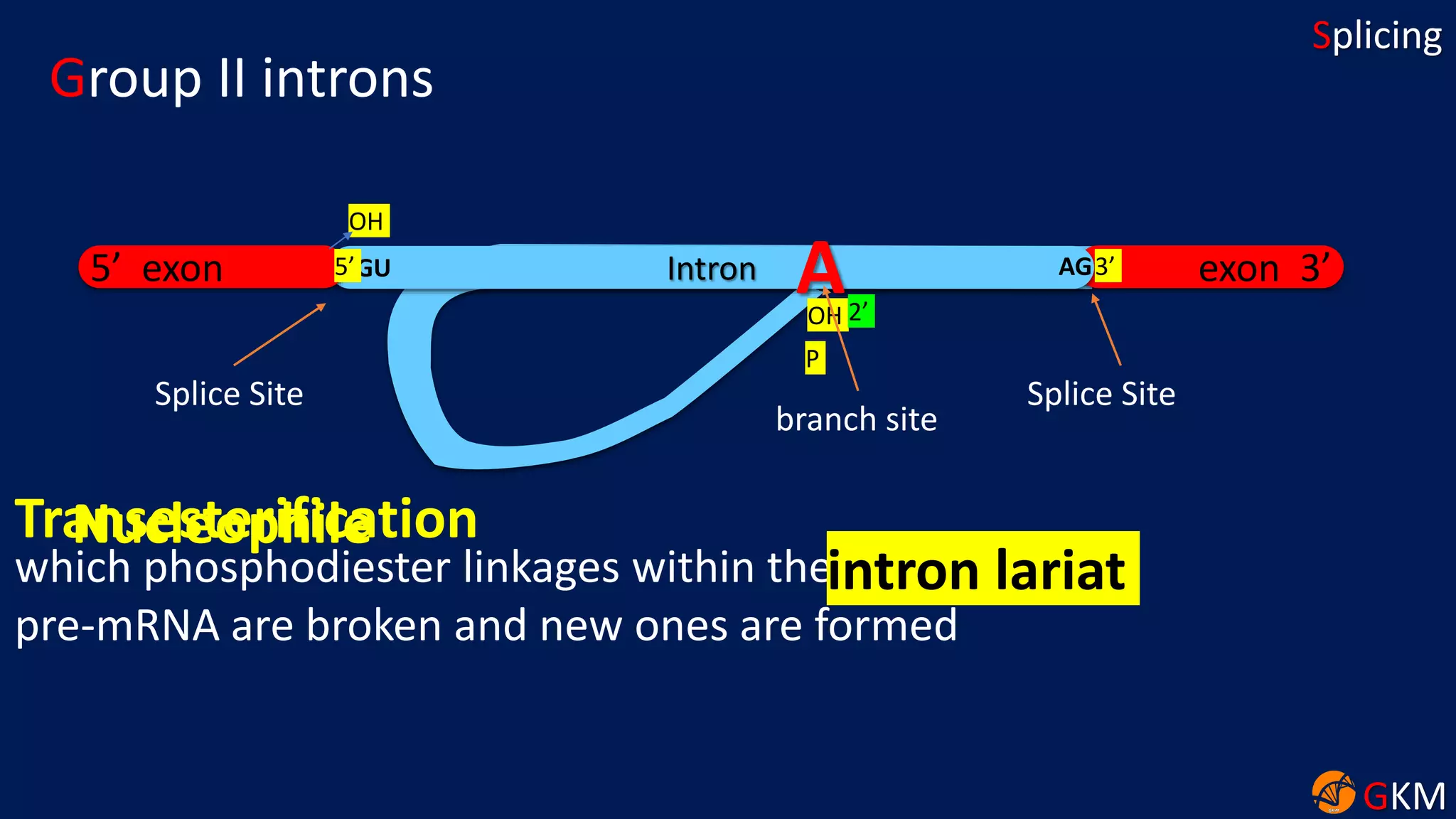
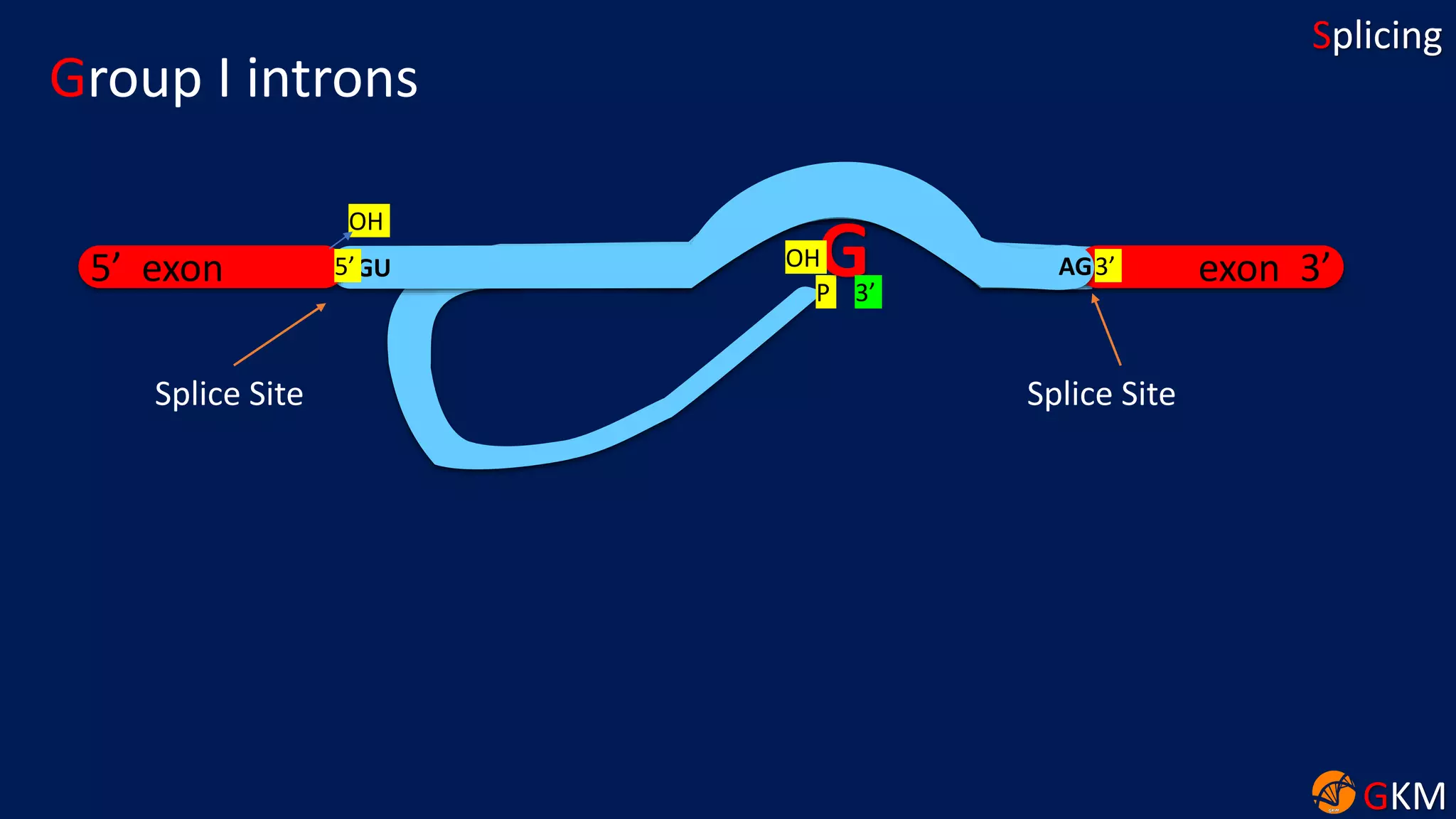
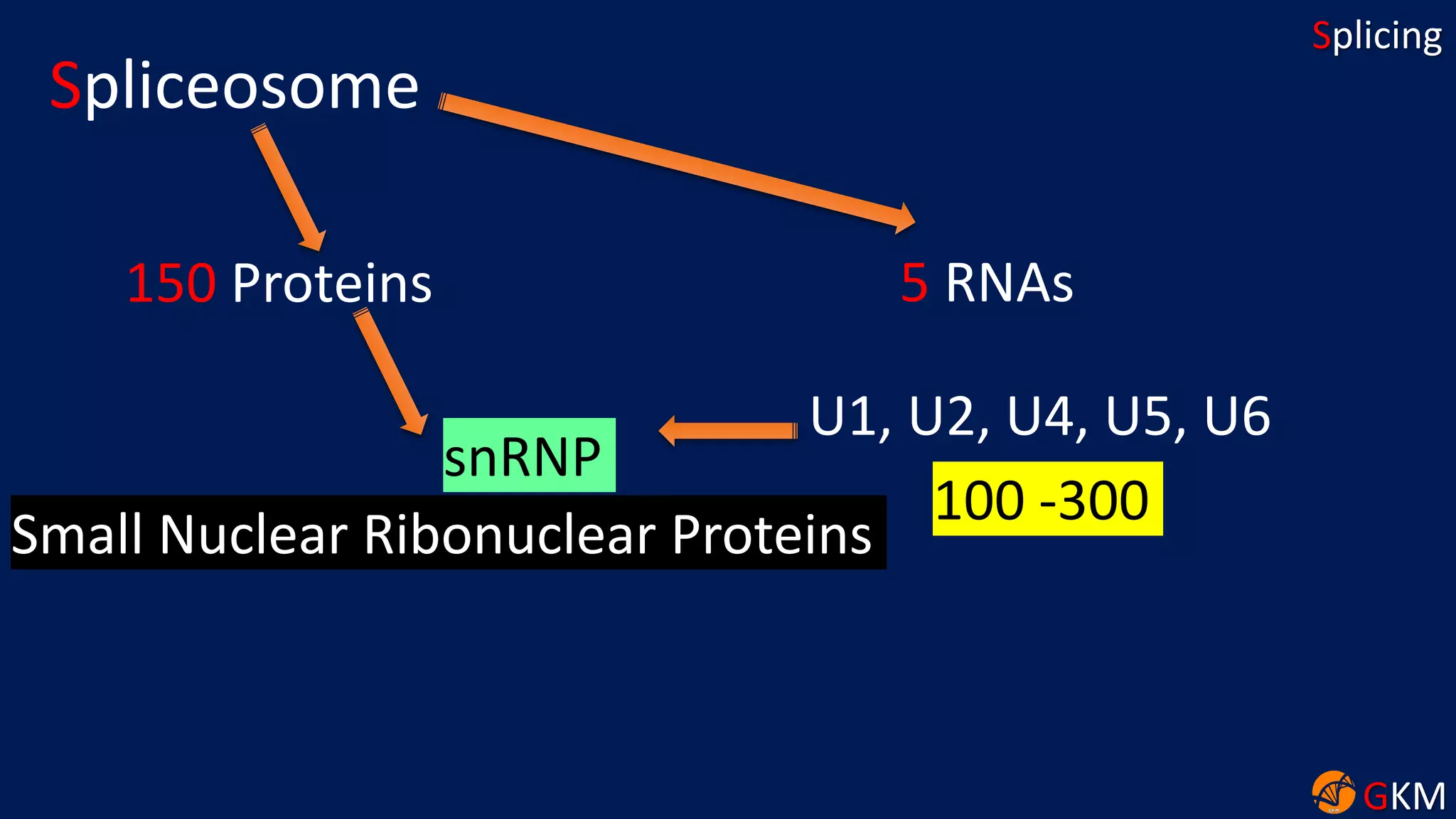
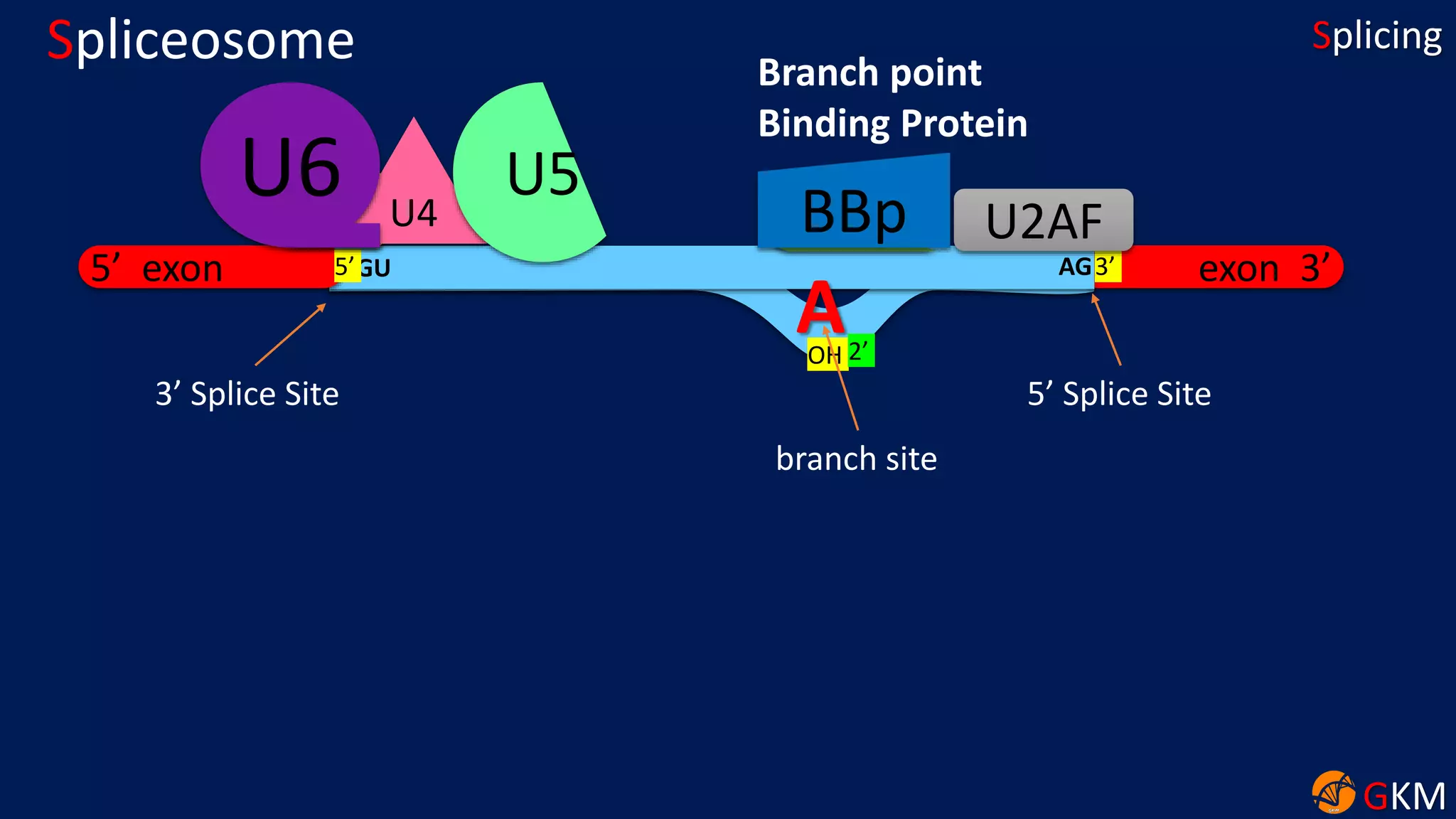
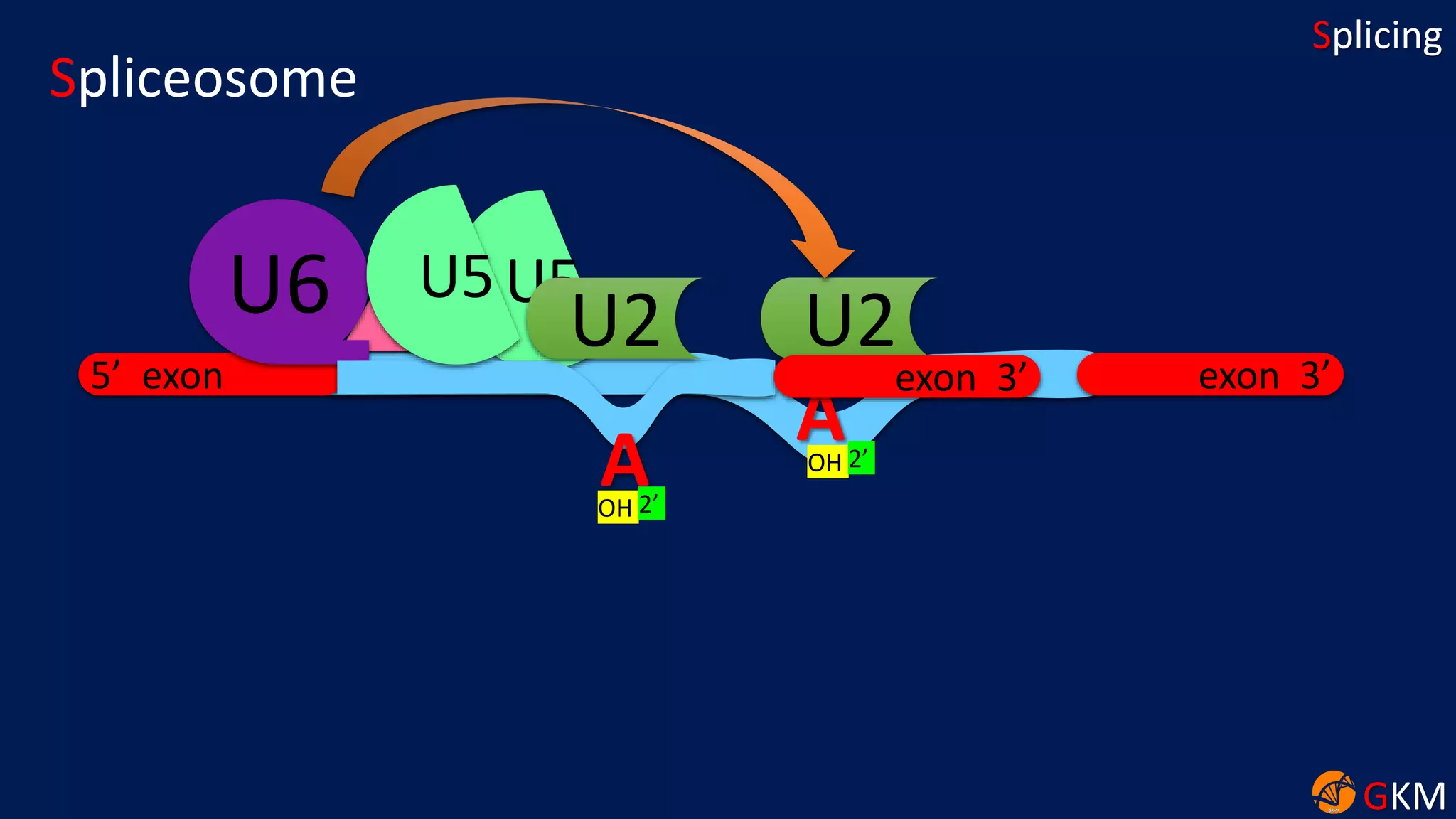
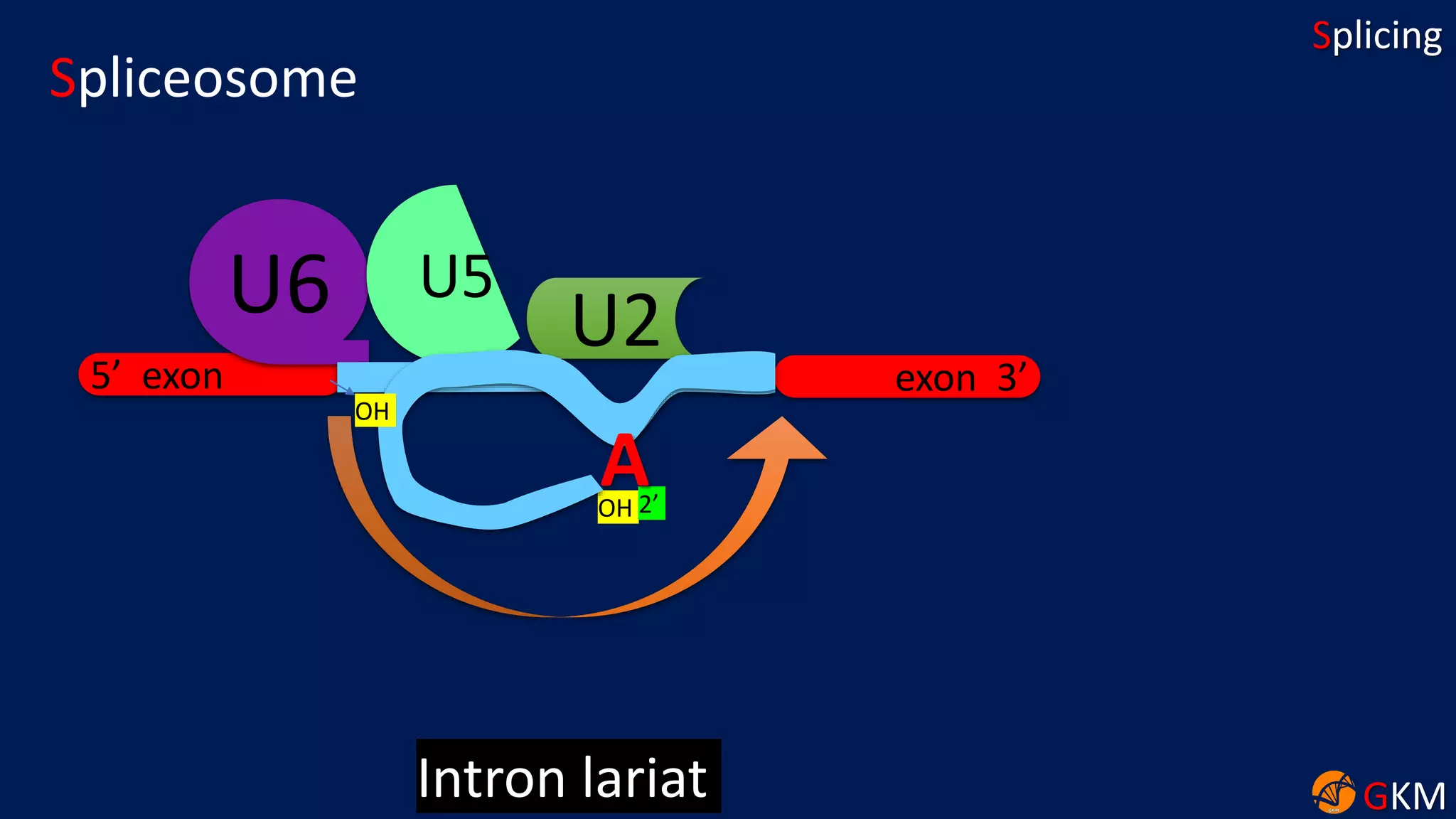

The document discusses RNA splicing, focusing on the removal of introns and joining of exons through processes involving different classes of introns (group I and II) and the spliceosome complex. It highlights the roles of various RNA and protein components, including snRNPs, in the splicing mechanism which involves transesterification and the formation of intron lariats. Key splice sites and binding proteins are also mentioned in relation to the splicing process.







