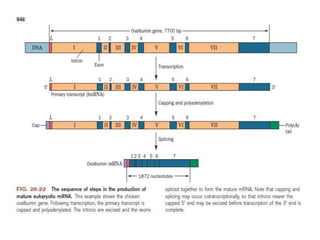
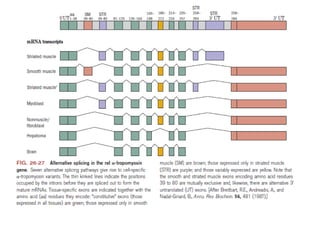
1) Eukaryotic mRNAs undergo post-transcriptional processing in the cell nucleus, including addition of a 5' cap and 3' polyadenylated tail.
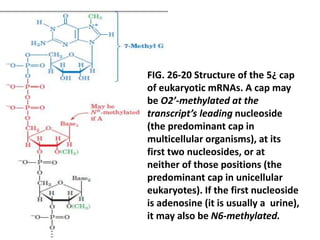
2) The 5' cap consists of a 7-methylguanosine residue joined to the initial nucleotide via a 5'-5' triphosphate bridge, and is added co-transcriptionally.
3) Mature mRNAs also contain a poly(A) tail of around 250 nucleotides added by poly(A) polymerase to the 3' end after cleavage of the primary transcript.