Embed presentation
Download to read offline
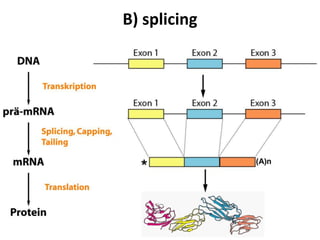
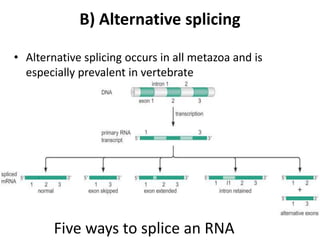

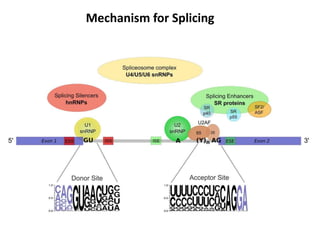

There are five ways that RNA can be spliced in eukaryotic cells. Splicing requires four intronic motifs at the 5' and 3' ends of introns. The spliceosome is composed of small nuclear RNAs and polypeptides that form snRNP complexes like U1, U2, U4, U5 and U6. Splicing is also directed by regulators like SR proteins that recognize enhancer sequences and hnRNPs that recognize silencer sequences to influence splicing.




