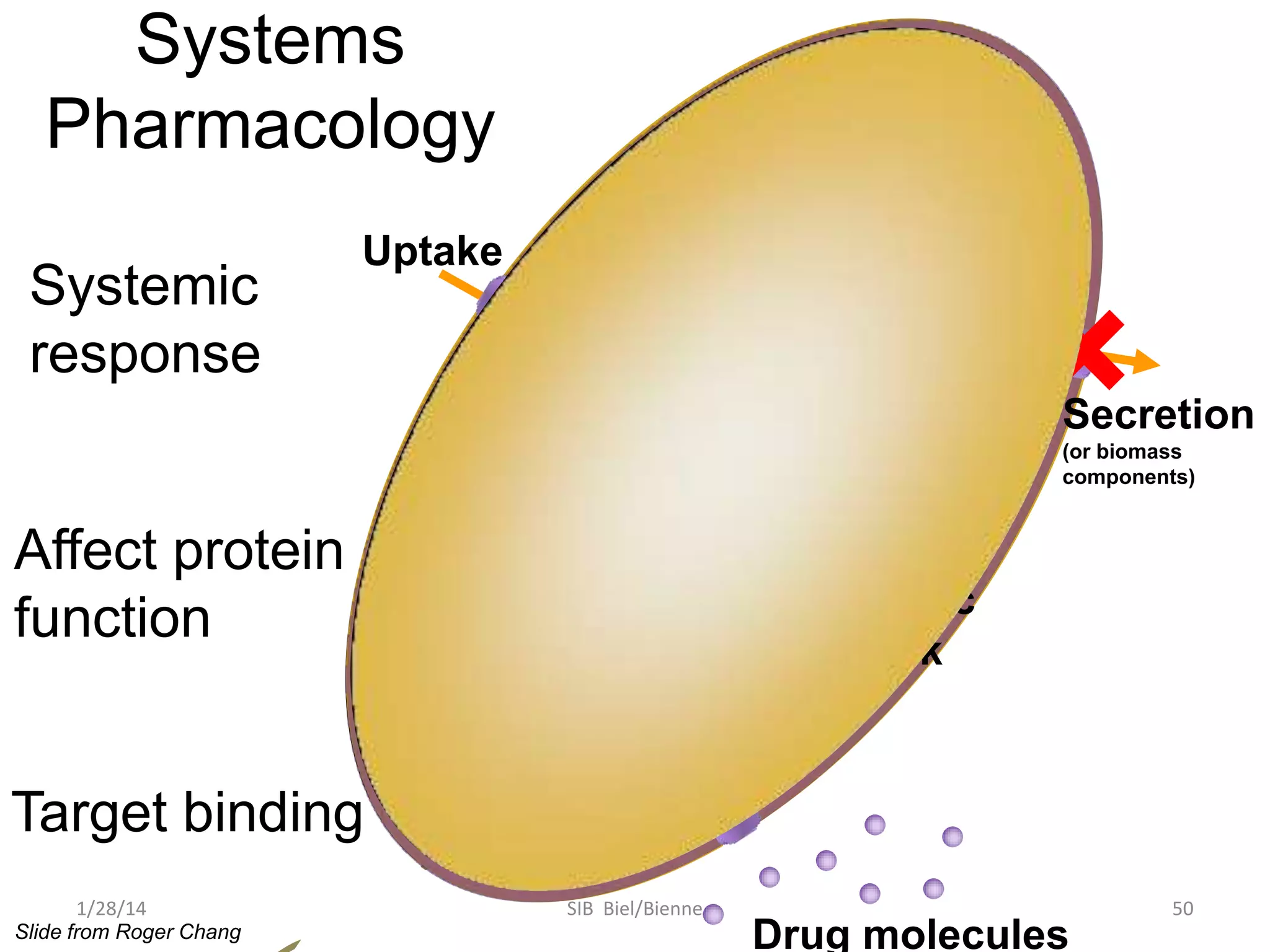
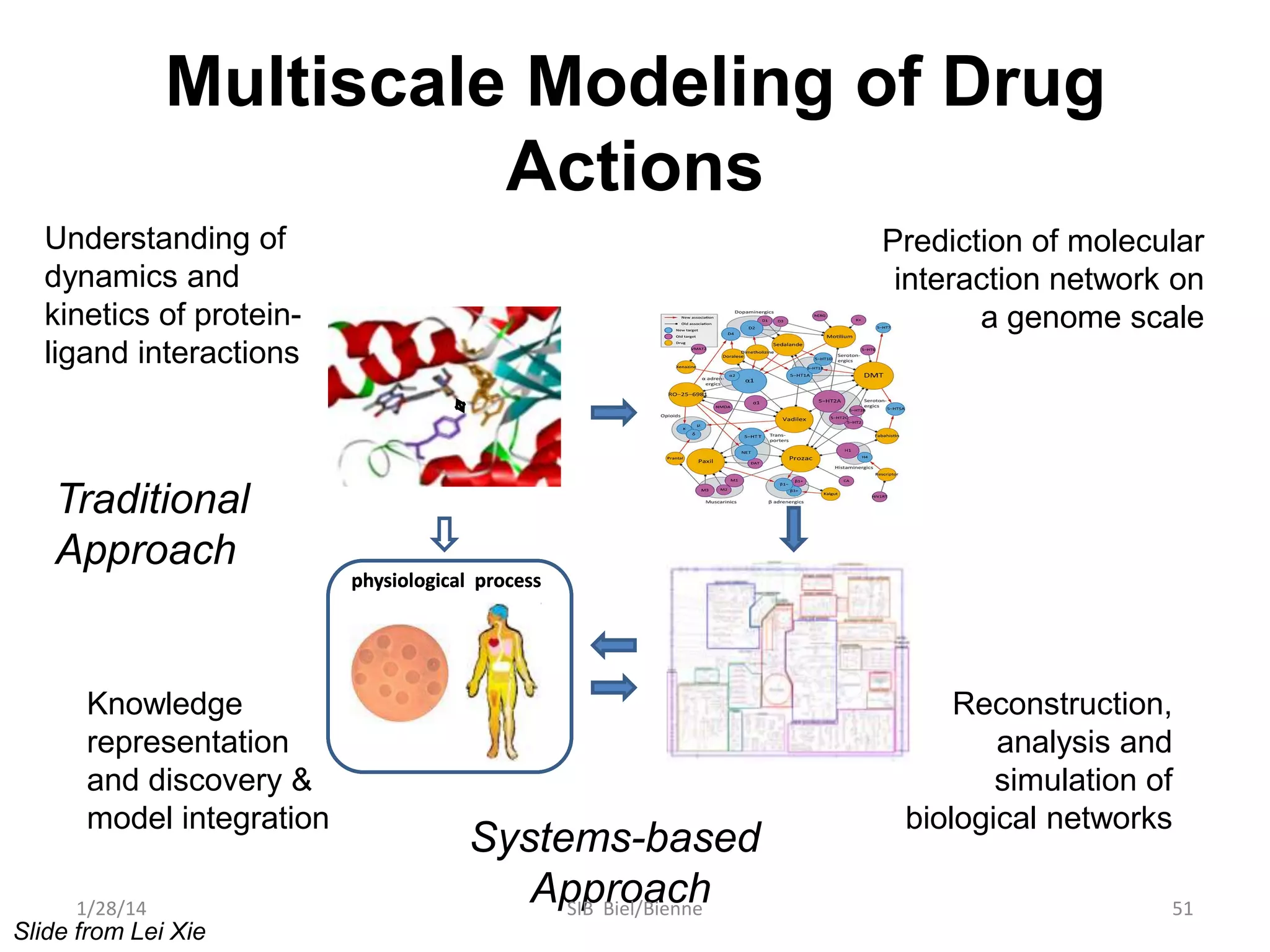
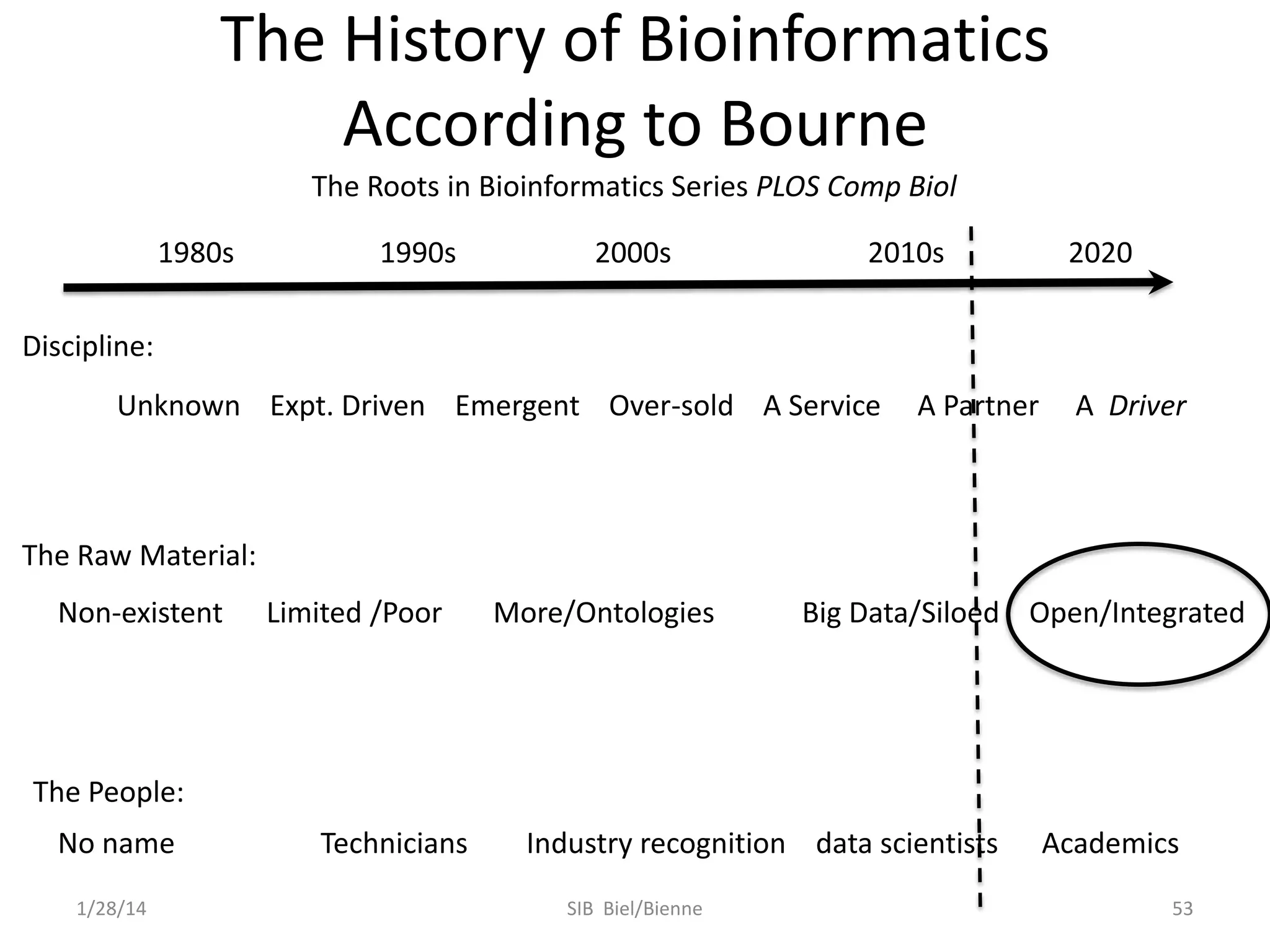
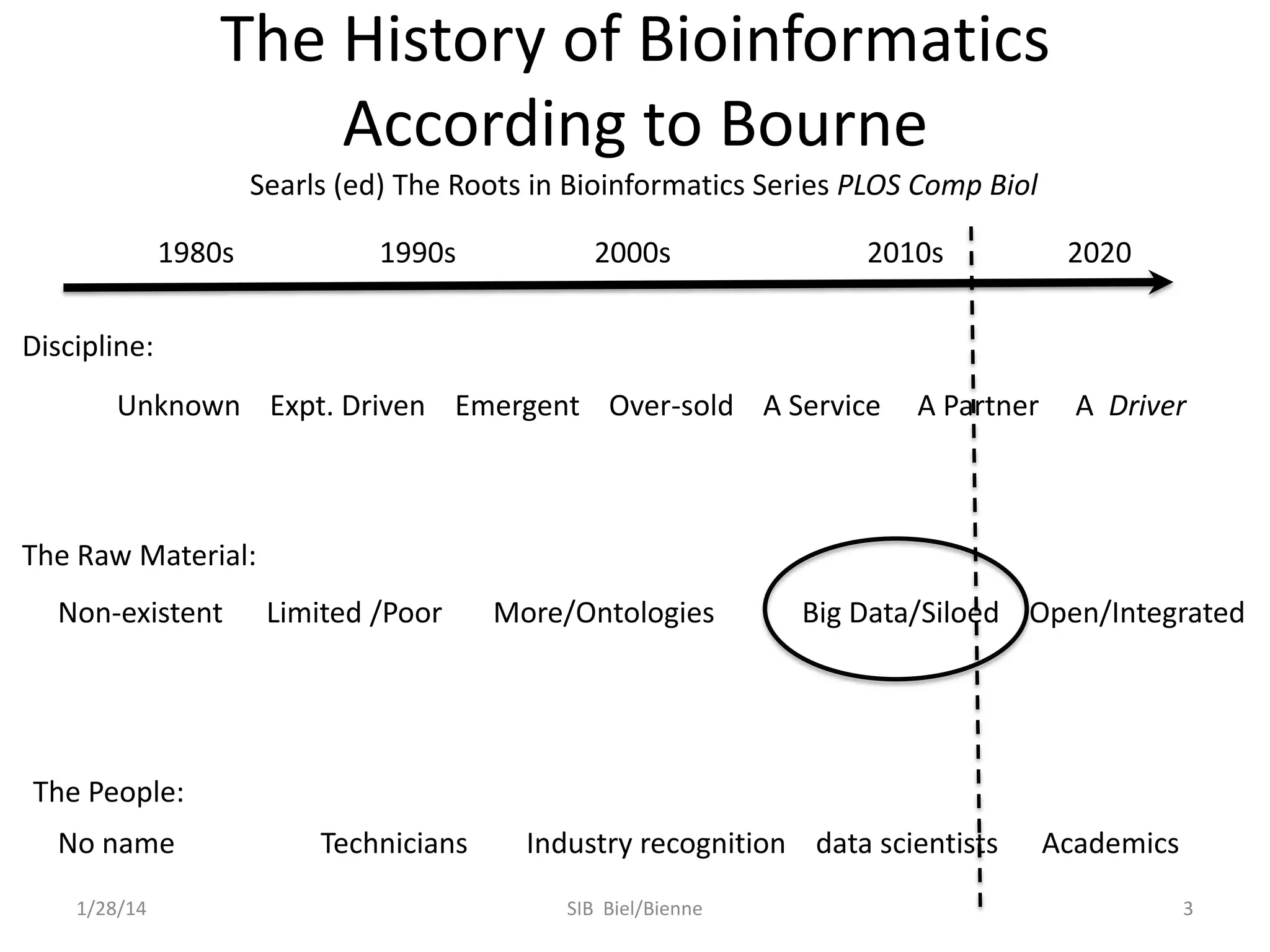
Philip E. Bourne discusses the evolution of bioinformatics in the context of open science and big data, emphasizing the importance of data quality, sustainability, and the need for collaboration across various disciplines and boundaries. He outlines the transition from a fragmented research lifecycle to a more integrated digital enterprise, where data and information can be accessed and utilized more effectively. The document also highlights the role of the NIH in facilitating open data practices and advancing data science initiatives.



![Web Logs etc. Are
Not Enough
Structure Summary page activity for
H1N1 Influenza related structures
Jan. 2008
Jul. 2008
Jan. 2009
Jul. 2009
Jan. 2010
Jul. 2010
3B7E: Neuraminidase of A/Brevig Mission/1/1918
H1N1 strain in complex with zanamivir
1RUZ: 1918 H1 Hemagglutinin
1/28/14
5
* http://www.cdc.gov/h1n1flu/estimates/April_March_13.htm
SIB Biel/Bienne
[Andreas Prlic]](https://image.slidesharecdn.com/sib0114-140128023905-phpapp01/75/Bioinformatics-in-the-Era-of-Open-Science-and-Big-Data-5-2048.jpg)



























![automate: workflows, pipeline &
service integrative frameworks
CS
SE
pool, share & collaborate web
systems
scientific software
engineering
semantics & ontologies
machine readable documentation
nanopub
1/28/14
[Carole Goble]
SIB Biel/Bienne
38](https://image.slidesharecdn.com/sib0114-140128023905-phpapp01/75/Bioinformatics-in-the-Era-of-Open-Science-and-Big-Data-38-2048.jpg)

![http://sagecongress.org/Presentations/Sommer.pdf
[Josh Sommer]
1/28/14
SIB Biel/Bienne
40](https://image.slidesharecdn.com/sib0114-140128023905-phpapp01/75/Bioinformatics-in-the-Era-of-Open-Science-and-Big-Data-40-2048.jpg)
![http://sagecongress.org/Presentations/Sommer.pdf
[Josh Sommer]
1/28/14
SIB Biel/Bienne
41](https://image.slidesharecdn.com/sib0114-140128023905-phpapp01/75/Bioinformatics-in-the-Era-of-Open-Science-and-Big-Data-41-2048.jpg)