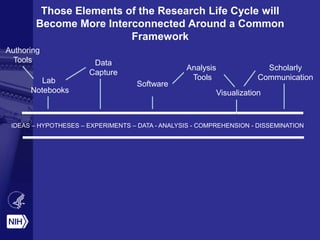
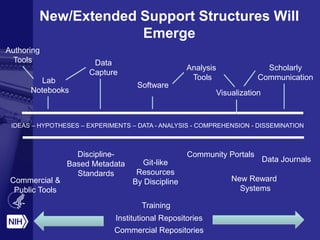
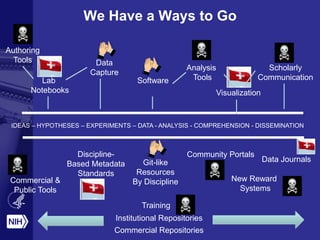
Philip E. Bourne discusses the challenges and opportunities in biomedical informatics and data sharing at the NIH, emphasizing the need for a sustainable data strategy and improved training in biomedical data science. He highlights the importance of understanding data usage and developing innovative funding models for data sustainability. The vision for a digital enterprise is portrayed as a means to enhance collaboration and efficiency in research through interconnected digital assets.



![Numberofreleasedentries
Year
Why Am I Here?
PDB Growth in Numbers and Complexity
[From the RCSB Protein Data Bank]](https://image.slidesharecdn.com/phrma-140423152256-phpapp02/85/PhRMA-Some-Early-Thoughts-4-320.jpg)








![* http://www.cdc.gov/h1n1flu/estimates/April_March_13.htm
Jan. 2008 Jan. 2009 Jan. 2010Jul. 2009Jul. 2008 Jul. 2010
1RUZ: 1918 H1 Hemagglutinin
Structure Summary page activity for
H1N1 Influenza related structures
3B7E: Neuraminidase of A/Brevig Mission/1/1918
H1N1 strain in complex with zanamivir
[Andreas Prlic]
Consider What
Might Be Possible](https://image.slidesharecdn.com/phrma-140423152256-phpapp02/85/PhRMA-Some-Early-Thoughts-13-320.jpg)










![47/53 “landmark” publications
could not be replicated
[Begley, Ellis Nature,
483, 2012] [Carole Goble]](https://image.slidesharecdn.com/phrma-140423152256-phpapp02/85/PhRMA-Some-Early-Thoughts-24-320.jpg)