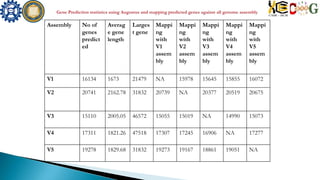
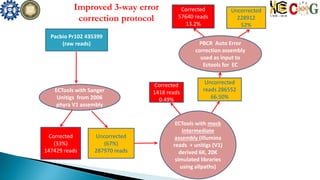
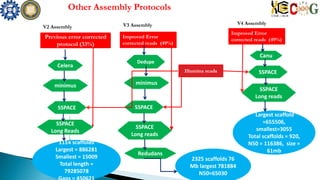
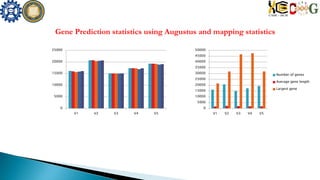
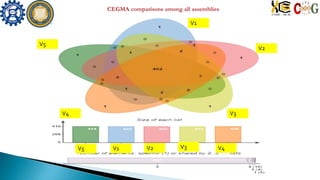
This document summarizes the assembly of the Phytophthora ramorum genome using PacBio long reads. It describes the error correction and assembly process for two P. ramorum strains, Pr102 and ND886. For Pr102, multiple assembly versions (V1-V5) were generated using different error correction and assembly protocols. The V5 assembly resulted in fewer scaffolds, larger size, and fewer gaps compared to previous versions. For ND886, PacBio reads were error corrected and assembled. Both assemblies captured more repetitive elements compared to previous Sanger-based assemblies. Gene predictions were also improved in number and quality.


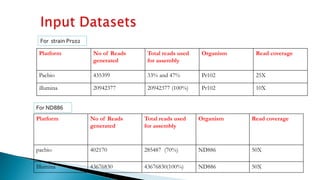


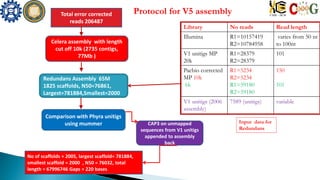
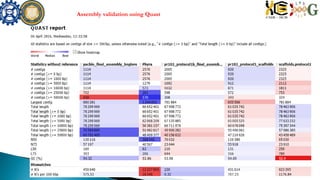



![ND886 assemblyTotal error corrected reads
285487
Celera Assembly
Minimus
Dedupe
Library No reads Read
length
Illumina R1=28389
986
R2=
28334221
varies
from 50 nt
to 100nt
Pacbio
corrected
MP 10k
6k
R1=
91555
R2=
91555
R1=13170
3
R2=13170
3
101
101
Read statistics
SSPACE [with illumina reads],Total contigs = 6443
Largest contig =648889,Smallest contig =2098,
assembly size = 150 Mb
Redudans
No of scaffolds = 2225, largest = 648906 ,
smallest = 2745 , N50 = 48161 , total length =
92877686 , Gaps = 4133
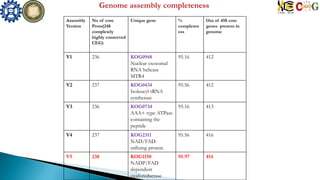
Assembly No of core
proteins
from 248
%
completen
ess
No of core
genes out
of 458
Nd886 234 94.35 410](https://image.slidesharecdn.com/ramorum2016final-160616100213/85/Ramorum2016-final-19-320.jpg)
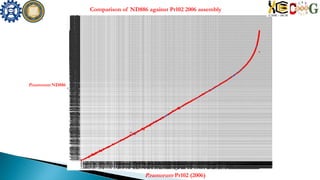


![Assemb
ly name
base
s
mask
ed
Sma
ll
RN
A
Simple
repeats
Low
complex
ity
GC
content
Total
interspe
rsed
repeats
LINE
[R2/R4/
NeSL]
Ty1/copio
Gypsy/DIRS1
LTR elements
DNA
transpos
on
Piggy
BAC
Tourist/
harbinge
r
V1 11.77
%
0.01
%
0.36% 0.03% 53.86% 11.37% 0.13% 10.01 % 1.23 % 0.16% 0.01%
V2 20.68
%
0.00
%
0.44 % 0.04 % 54.32% 20.20 % 0.32 % 5.64 % 1.48 % 0.16 % 0.0.1 %
V3 27.00
%
0.45
%
0.75% 0.12% 52.40 % 25.70 % 0.39 % 23.85 % 1.46 % 0.17 % 0.01 %
V4 21.06
%
0.11
%
0.53% 0.05% 54.09 % 20.37 % 0.29 % 18.70 % 1.38 % 0.17 % 0.0.1 %
V5 24.34
%
0.07
%
0.49% 0.06 % 53.98 % 23.73% 0.37% 21.89 % 1.47 % 0.15 % 0.0.1%
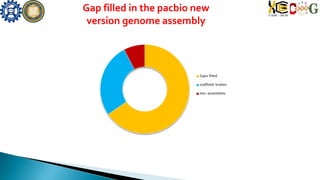
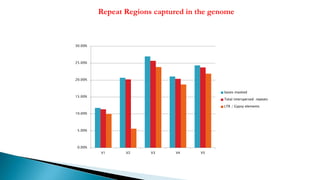
Repeat Regions captured in the genome](https://image.slidesharecdn.com/ramorum2016final-160616100213/85/Ramorum2016-final-23-320.jpg)

![Assembly
name
bases
masked
Small
RNA
Simple
repeats
Low
complexity
GC
content
Total
interspersed
repeats
LINE
[R2/R4/
NeSL]
Ty1/copio
Gypsy/DIRS1
LTR elements
DNA transposon Piggy BAC Tourist/harbinger
P.ramorum
2006
7847064 bp
(11.77%)
11 (6033
bp)
0.01%
5336 (242077
bp)
0.36%
422(20747 bp)
0.03%
53.86% 7580618 bp
(11.37%)
53 (88470
bp 0.13%)
5972 (6669143
bp) 10.01 %
1174 (823005bp)
1.23 %
200 (104977 bp)
0.16%
12 (5609 bp)
0.01%
Protocol
1b
16553511
bp
(24.34%)
75 (49453
bp)
0.07%
7122(331100
bp)
0.49%
816 (40373
bp)
0.06 %
53.98 % 16138229 bp
(23.73%)
87
(250632bp
)
0.37%
8822(14885437
bp ) 21.89 %
1419 ( 1002160 bp )
1.47 %
198 ( 104684 bp )
0.15 %
13 (5809 bp )
0.0.1%
Protocol 2 21185972
bp
(27.00%)
605(3493
28 bp)
0.45%
11702
(586604 bp)
0.75%
1787 (91389
bp)
0.12%
52.40 % 20163370 bp
25.70 %
112
308607 bp
0.39 %
11127 (
18710327 bp )
23.85 %
1756
(1144436) bp
1.46 %
231
129697 bp
0.17 %
12
5417 bp
0.01 %
Protocol 3 12854764
bp
(21.06 %)
64( 69255
bp)
0.11 %
6801 (323105
bp)
0.53%
679 (33221
bp)
0.05%
54.09 % 12434182 bp
20.37 %
64 (176881
bp)
0.29 %
6752
(11415393 bp )
18.70 %
1133 (841908 bp) 1.38
%
191 (105824 bp)
0.17 %
12 (6211 bp)
0.0.1 %
Bangalore
meeting
16192690
bp
(20.68 %)
8 (3317
bp)
0.00%
7549 (340933
bp)
0.44 %
699(33372 bp)
0.04 %
54.32% 15819353 bp
(20.20 %)
92
250092bp
(0.32 %)
2498 (4413567
bp) 5.64 %
1560 (1155118 bp )
1.48 %
228 (126831bp)
0.16 %
15 (6376 bp)
0.0.1 %
Repeat Regions captured in the genome](https://image.slidesharecdn.com/ramorum2016final-160616100213/85/Ramorum2016-final-25-320.jpg)