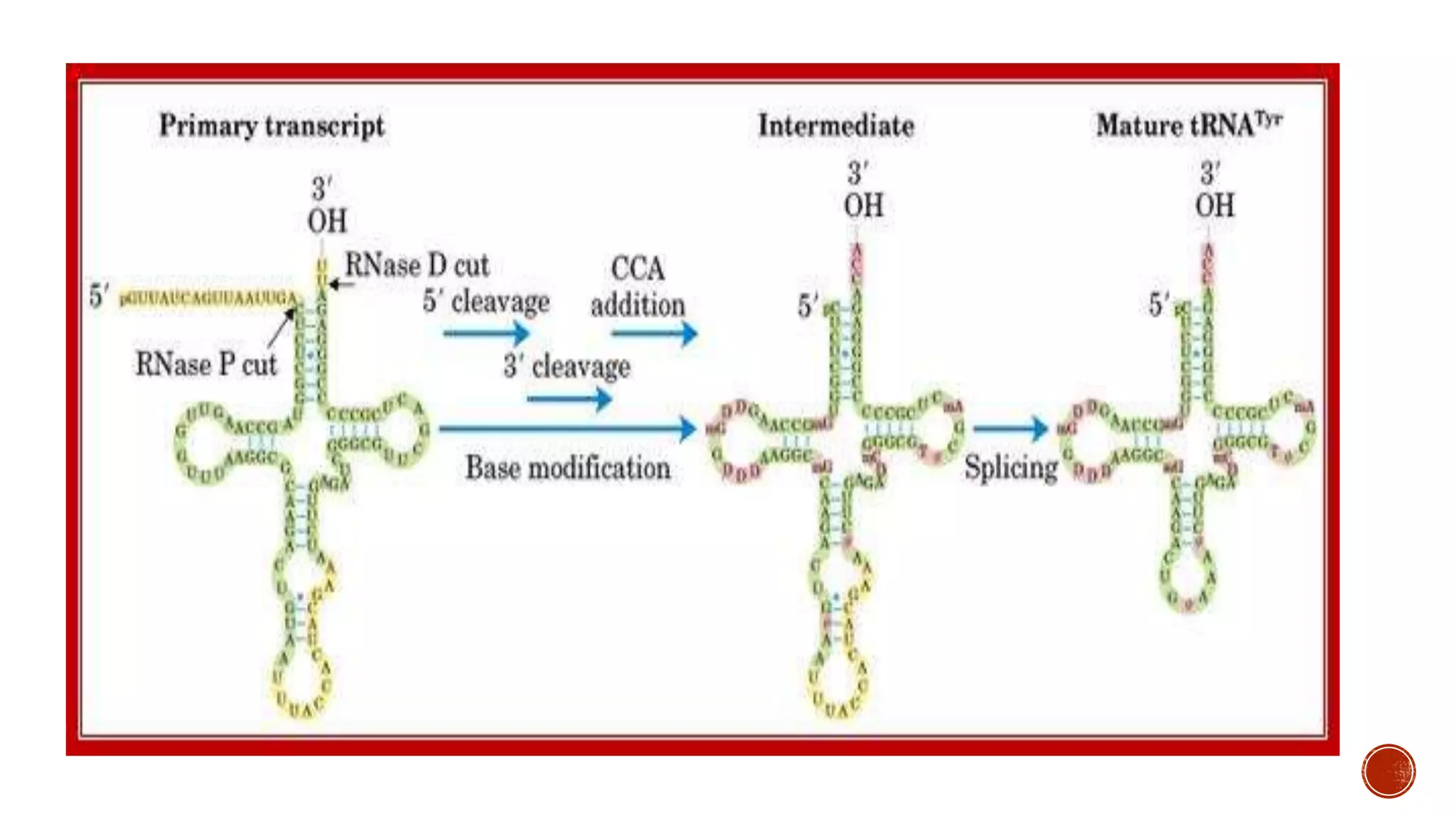
The document summarizes the processing of transfer RNA (tRNA) in organisms. TRNA undergoes extensive processing where nucleotides are removed from both the 5' and 3' ends by endonucleases and exonucleases to generate mature tRNA that is 80-90 nucleotides long. A key step is the addition of the 3' terminal CCA sequence by tRNA nucleotidyl transferase. The mature tRNA also contains various modified bases introduced by enzymatic modifications. Introns are also excised from precursor tRNA to form mature functional tRNA for protein synthesis.