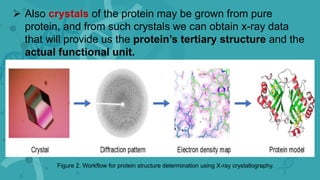
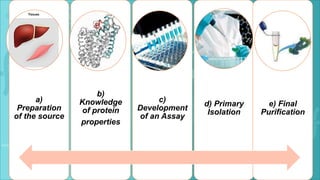
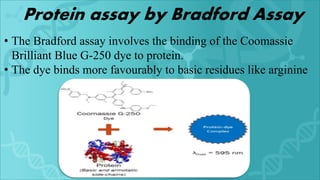
Protein purification is a series of processes to isolate a protein from a complex mixture. It is important to characterize a protein's function, structure, and interactions. The general steps of purification include preparing the source, exploiting differences in protein properties through separation techniques, and monitoring purity with assays. Purification yields a pure protein sample for determining its sequence, structure, and role. Affinity tags and other tags are often used to aid purification. Biopharmaceuticals are medically important proteins produced through biotechnology.