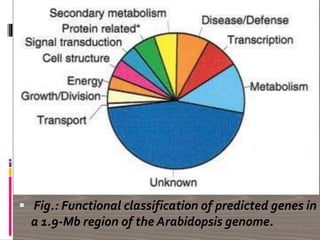
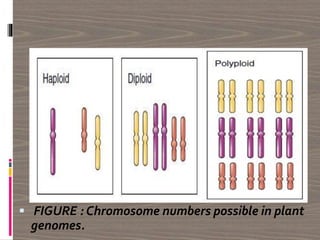
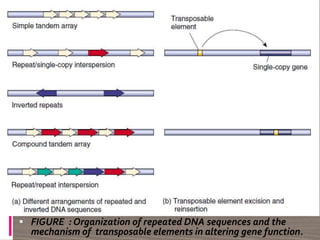
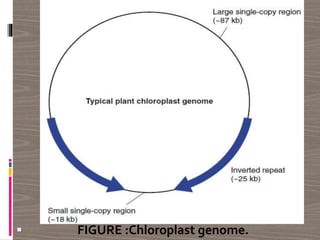
The document discusses plant genomics, covering the history, study, and organization of plant genomes, including the significance of model organisms like Arabidopsis thaliana. It highlights the complexity and variability of plant genomes, the roles of chloroplast and mitochondrial DNA, and the impact of transposable elements. Additionally, it emphasizes the applications of genomics in improving crop productivity and quality, showcasing its relevance in food production and environmental sustainability.