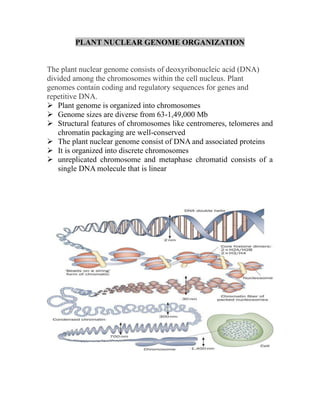
Plant nuclear genome organization
- 1. PLANT NUCLEAR GENOME ORGANIZATION The plant nuclear genome consists of deoxyribonucleic acid (DNA) divided among the chromosomes within the cell nucleus. Plant genomes contain coding and regulatory sequences for genes and repetitive DNA. ➢ Plant genome is organized into chromosomes ➢ Genome sizes are diverse from 63-1,49,000 Mb ➢ Structural features of chromosomes like centromeres, telomeres and chromatin packaging are well-conserved ➢ The plant nuclear genome consist of DNA and associated proteins ➢ It is organized into discrete chromosomes ➢ unreplicated chromosome and metaphase chromatid consists of a single DNA molecule that is linear
- 2. Proteins ➢ There are 2 types of proteins found in DNA Basic proteins-Positive charge at neutral pH Histone (H1, H2A, H2B, H3 and H4 ) ➢ Acidic proteins- Negative charge at neutral pH Non-histone proteins DNA PACKING: ➢ Nucleosome contains Eight histone protein molecules, these being two each of histones H2A, H2B, H3 and H4 ➢ These eight proteins form a barrel shaped core octamer with the DNA twice around the outside ➢ 140 to 150 bp of DNA are associated with the nucleosome particle and each nucleosome is separated by 50-70 bp of linker DNA constituent of chromatin ➢ DNA= 20-40 %- most important chemical constituent of chromatin ➢ RNA=05-10 %-associated with chromatin as; ⚫ Ribosomal RNA-( rRNA) ⚫ Messenger RNA- (mRNA) ⚫ Transfer RNA- (tRNA) ➢ PROTEINS=55-60%-associated with chromatin TYPES OF CHROMATIN
- 3. Euchromatin ➢ Lightly packed form of chromatin that is rich in gene concentration ➢ takes up light stain and represent most of the chromatin, that disperse after mitosis has completed. ➢ Consists of structural genes which replicate and transcribe during G1 and S phase of the interphase. ➢ Considered genetically active chromatin, since it has a role in their phenotypic expression of the genes. Heterochromatin ➢ Tightly packed form of chromatin that takes up deep stain during interphase and prophase but metaphase takes up light stain. ➢ Chromomeres, centromeric regions, and knobs also take up dark staining, of which centromeric regions and knobs are the true Heterochromatic. ➢ In the chromosomes all the centromeres fuse to form a long Heterochromatic mass called chromocentre. ➢ Heterochromatin consists of highly repetitive DNA sequences. CENTROMERE ➢ The centromere is the specialized DNA sequence of a chromosome that links a pair of sister chromatids ➢ During mitosis, spindle fibers attach to the centromere via the kinetochore.
- 4. TELOMERE ➢ sequences consist of short tandem repeats that contribute to the stability and integrity of the chromosome
- 5. NUCLEAR GENOME Single-or low-copy sequences ➢ Single copy DNA refers to any DNA sequence present in only one copy per haploid genome. ➢ It contain INTRON AND EXON ➢ The average coding region of plant gene requires about 1,300 bp or 15,000 genes.
- 6. Repetitive DNA-non coding Repeated sequences are patterns of nucleic acids (DNA) that occur in multiple copies throughout the genome Repetitive DNA can be present in hundreds or even thousands of copies in the genome Repetitive DNA can be subdivided into 2 classes • Tandem repeats • Dispersed repeats Tandem repeats ➢ Much of the repetitive DNA falls into the tandem repeats is referred to as satellite DNA ➢ It is a non-coding DNA seq ➢ Satellite DNA is
- 7. a ssociated with centromere or telomeres and is heterochromatic Moderately repetitive DNA includes: ➢ minisatellites ➢ Microsatellites ➢ variable number tandem repeats (VNTRs) Dispersed repeats ➢ Dispersed repeats are segments of DNA that occur multiple times at more or less random positions in the genome. ➢ They are typically transposable elements, large segments that encode a protein responsible for the moving of the segment from one site to another. Transposable elements ➢ DNA transposons ➢ Retrotransposons DNA transposons ➢ DNA transposons it also called JUMPING GENE. ➢ Eukaryotic genomes contain an abundance of repeated DNA, and some repeated sequences are mobile. ➢ Transposable elements (TEs) are defined as DNA sequences that are able to move from one location to another in the genome. ➢ It move from one genomic location to another by a cut-and-paste
- 8. mechanism. REFERENCE 1. https://www.researchgate.net/publication/228052081_Plant_Nuclear _Genome_Composition 2. Capelson M, Liang Y, Schulte R, Mair W, Wagner U, Hetzer MW (2010) Chromatin-bound nuclear pore components regulate gene expression in higher eukaryotes. 3. Alkhimova, O.G., Mazurok, N.A., Potapova, T.A., Zakian, S.M., Heslop‐Harrison, J.S. and Vershinin, A.V. (2004) Diverse patterns of the tandem repeats organization in rye chromosomes.