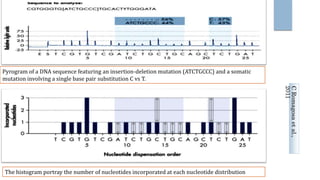
DNA sequencing determines the order of nucleotides in a DNA molecule. Next-generation sequencing (NGS) methods like pyrosequencing have accelerated research by allowing high-throughput, low-cost sequencing. Pyrosequencing works by detecting pyrophosphate release during DNA synthesis. It has applications in genetics, epigenetics, forensics, medicine, and more. NGS continues to advance sequencing capabilities and make whole genome analysis increasingly accessible.