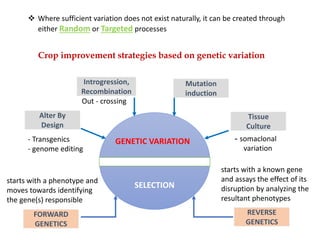
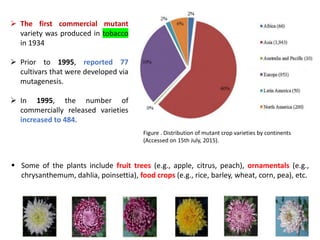
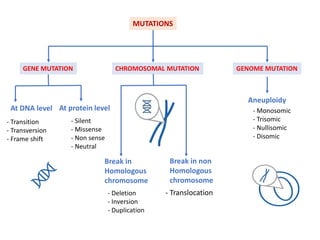
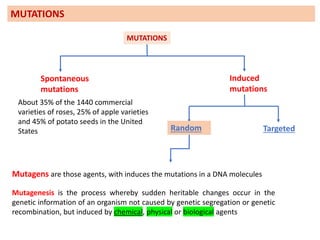
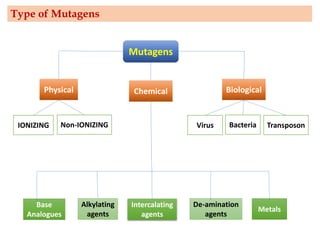
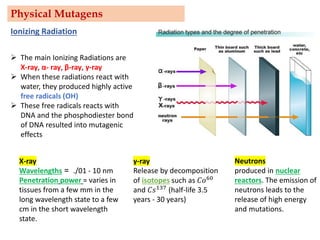
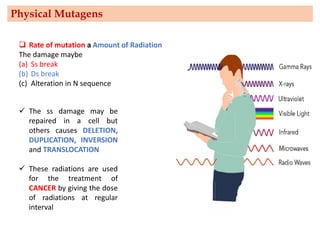
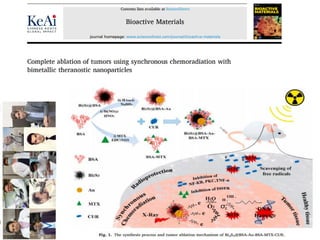
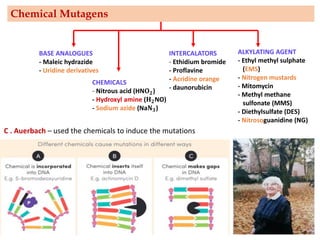
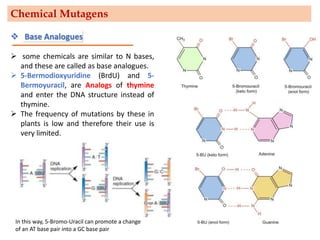
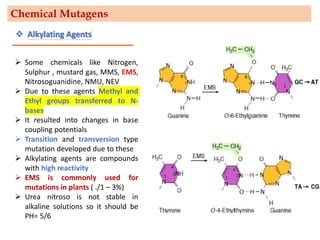
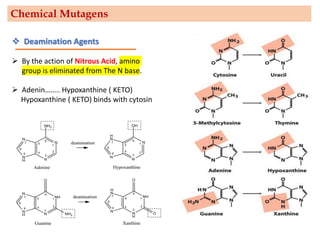
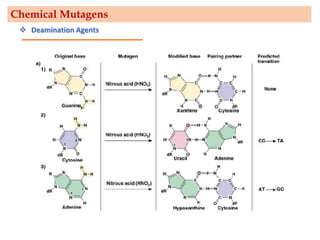
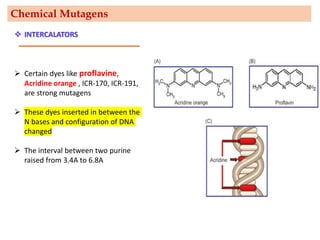
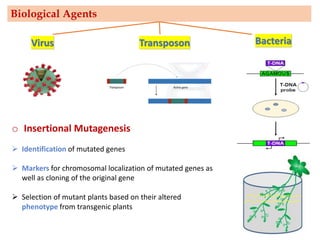
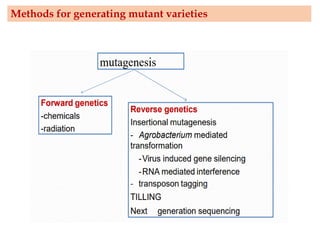
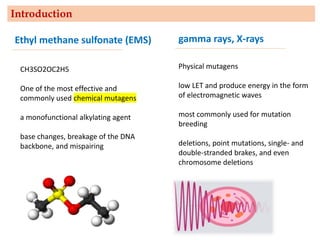
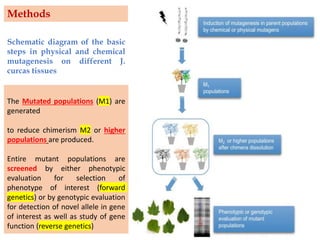
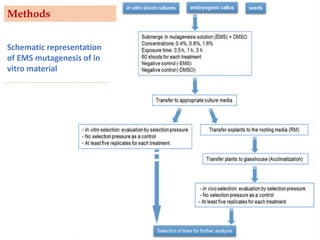
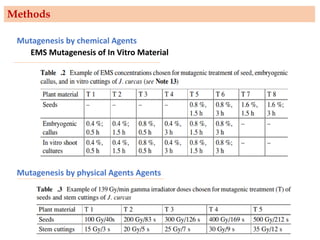
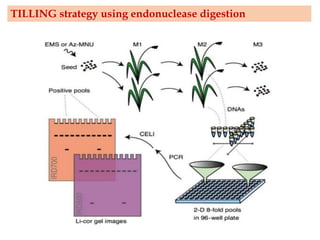
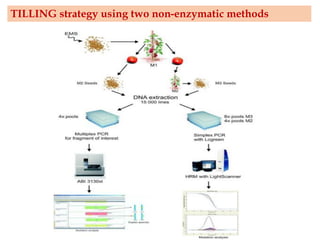
Mutagenesis techniques like chemical mutagenesis using ethyl methane sulfonate (EMS) and physical mutagenesis using gamma irradiation and X-rays can be used to induce mutations in Jatropha curcas, an economically important plant. The document describes methods for mutagenesis in different tissues of J. curcas, including seeds, under in vitro and in vivo conditions. It also discusses approaches for detecting mutations, such as TILLING, and analyzing mutant populations to select plants with desirable traits for further breeding.