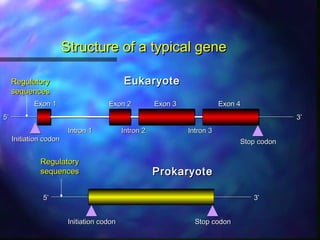
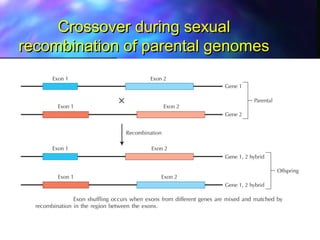
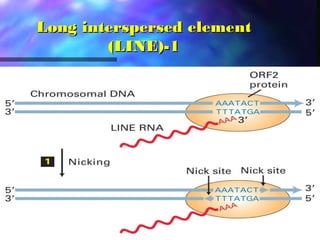
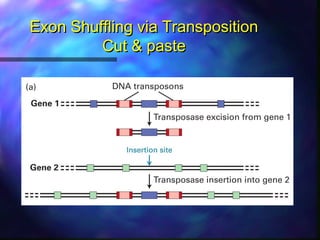
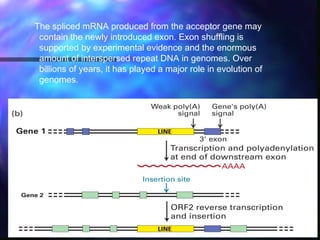
This document discusses exon shuffling, which is a mechanism by which new genes can form through the rearrangement of exons from different genes. Exon shuffling was first proposed in 1978 and involves recombination within introns that allows exons to be assorted independently, generating new exon combinations. There are three main types of exon shuffling: exon duplication, insertion, and deletion. Exon shuffling generates genetic variation and mosaic proteins, and it has played a major role in evolution. The mechanisms involved are crossover during sexual recombination and transposon-mediated movements that can cut, paste, or copy and paste exons into new locations.




















![References
https://www.google.com/search?
client=opera&q=scheme+of+presentation&sourceid=opera&ie=
UTF-8&oe=UTF-8#q=exon+shuffling
Wang, W., Zheng, H., Fan, C., Li, J., Shi, J., Cai, Z., . . . Wang,
J. (2006). High rate of chimeric gene origination by retroposition
in plant genomes. Plant Cell, 18(8), 1791-1802. doi:
tpc.106.041905 [pii] 10.1105/tpc.106.041905
Origins of New Genes: Exon
Shuffling
By Carl Hillstrom](https://image.slidesharecdn.com/exonshuffling-150522083244-lva1-app6891/85/Exon-shuffling-26-320.jpg)
