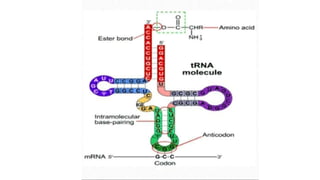
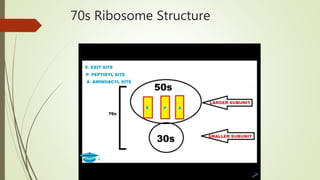
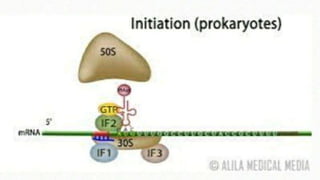
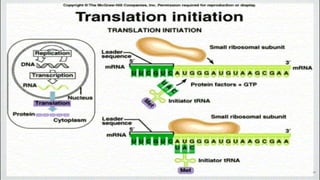
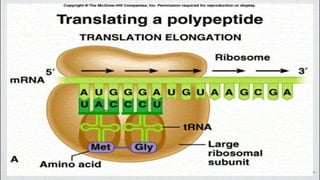
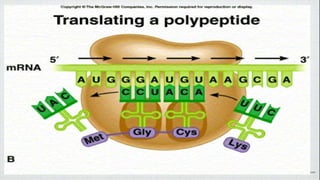
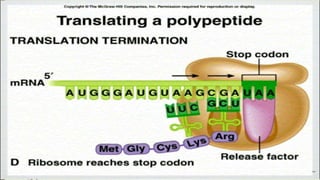
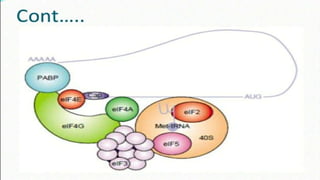
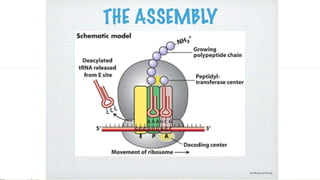
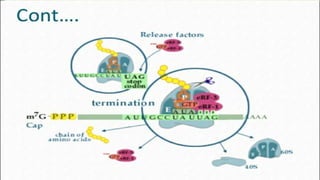
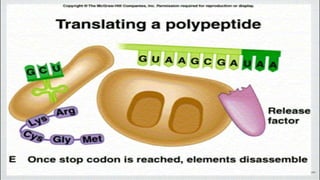
This document summarizes the process of translation in prokaryotes and eukaryotes. It discusses the central dogma of molecular biology and explains the four main steps of translation - initiation, elongation, termination, and activation. In prokaryotes, initiation requires initiation factors to form the 70S initiation complex, elongation is cyclic and uses elongation factors, and termination uses release factors. Eukaryotic translation is more complex, with initiation forming the 43S and 48S preinitiation complexes before joining the 60S subunit. Elongation and termination are similar to prokaryotes but use eukaryotic specific factors. Polyribosomes with multiple ribosomes on a single mRNA can increase protein production efficiency.