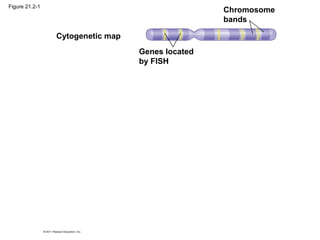
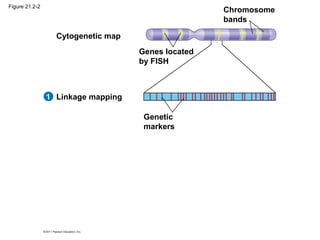
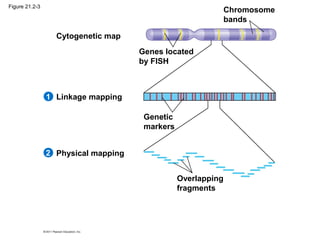
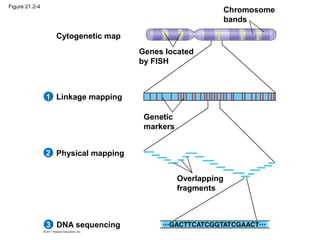
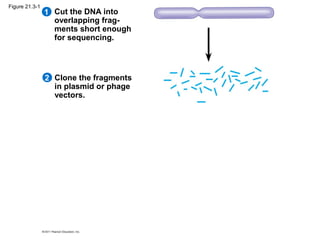
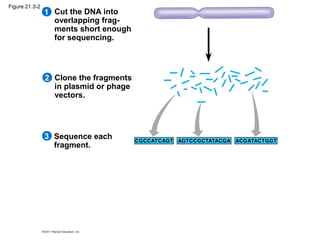
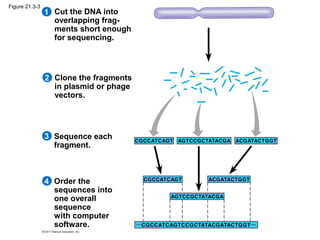
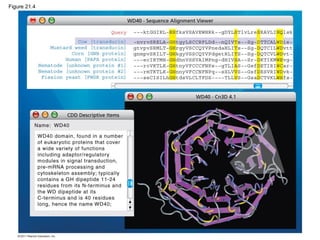
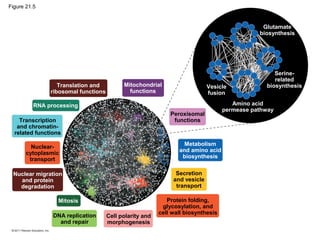
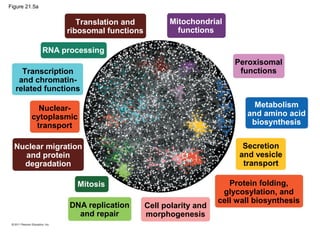
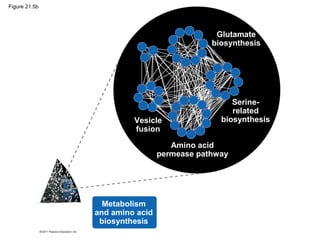
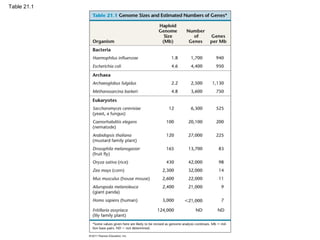
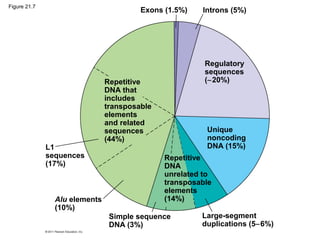
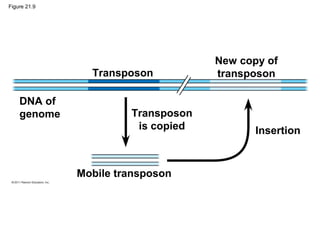
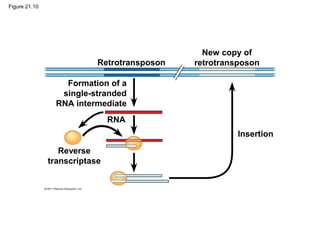
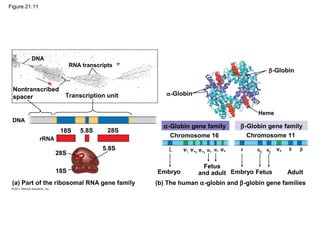
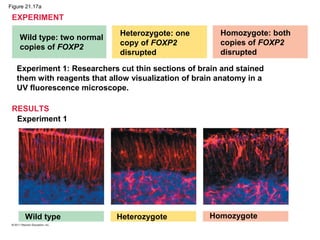
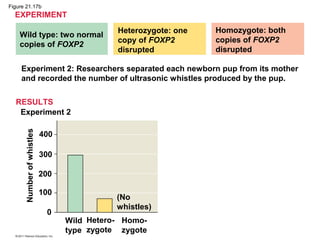
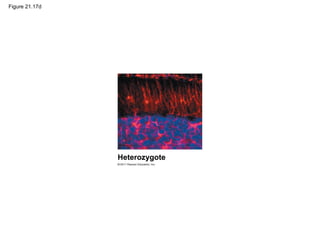
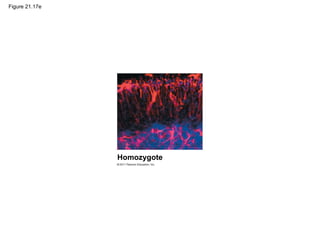
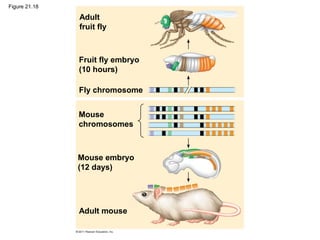
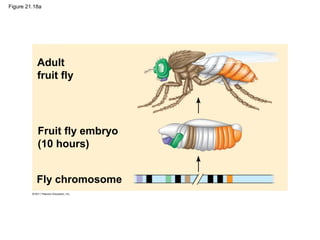
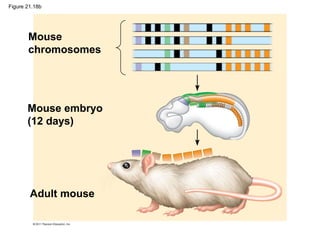
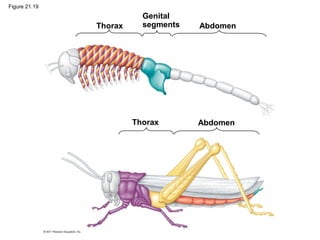
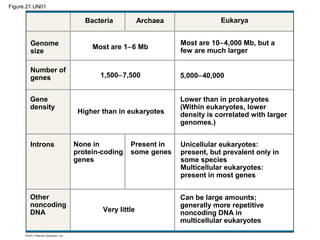
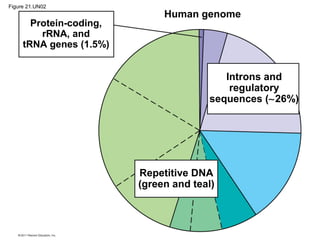
This document provides an overview of key concepts from Chapter 21 of Campbell Biology about genomes and their evolution. It discusses the stages of genome sequencing, including genetic mapping, physical mapping, and DNA sequencing. It also describes how new sequencing techniques like whole-genome shotgun sequencing have accelerated the process. Bioinformatics tools are now used to analyze whole genomes and understand gene functions on a systems level. Comparisons of complete genome sequences from different organisms reveal variation in genome size, number of genes, and amount of noncoding DNA between domains of life.