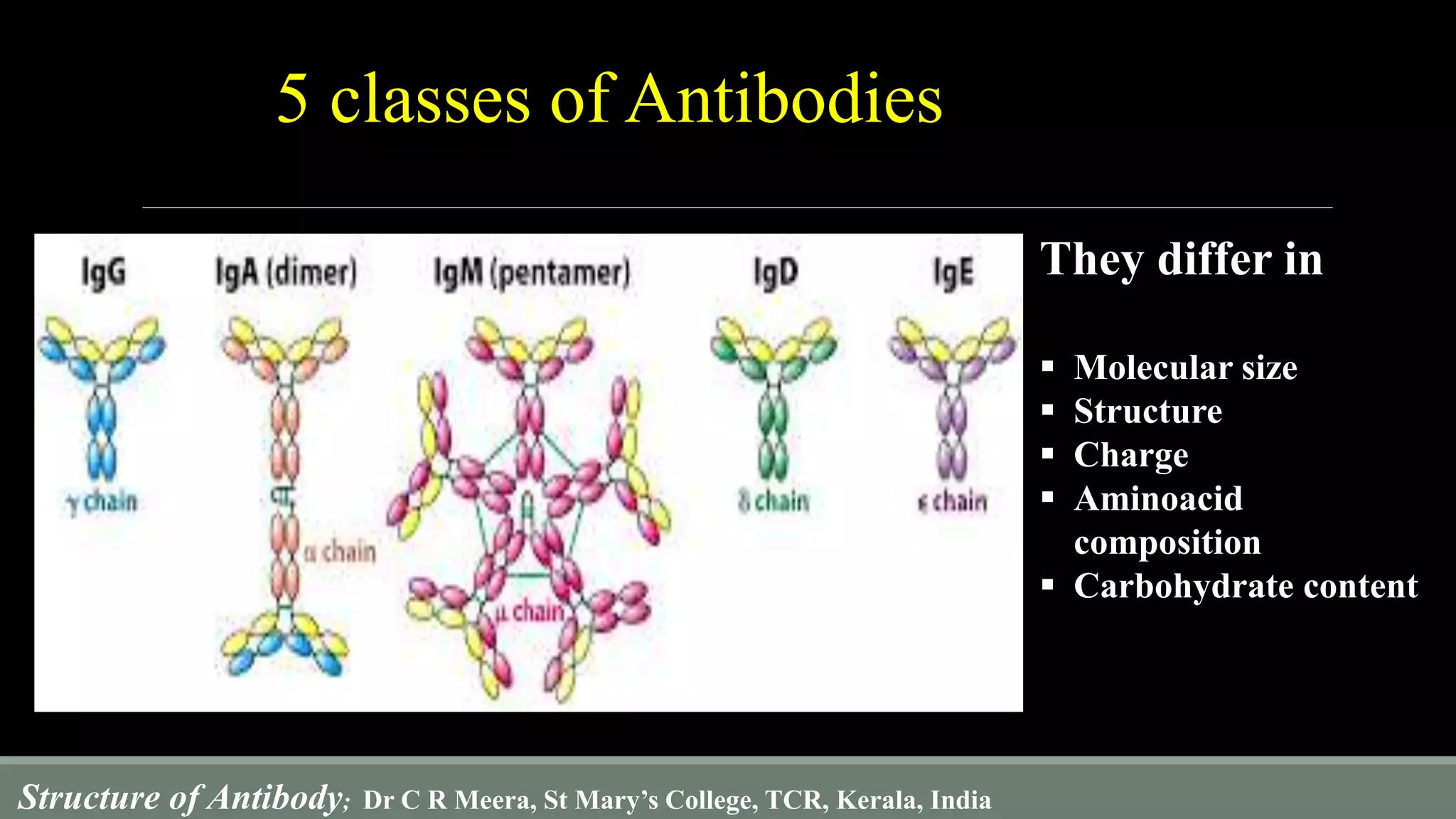
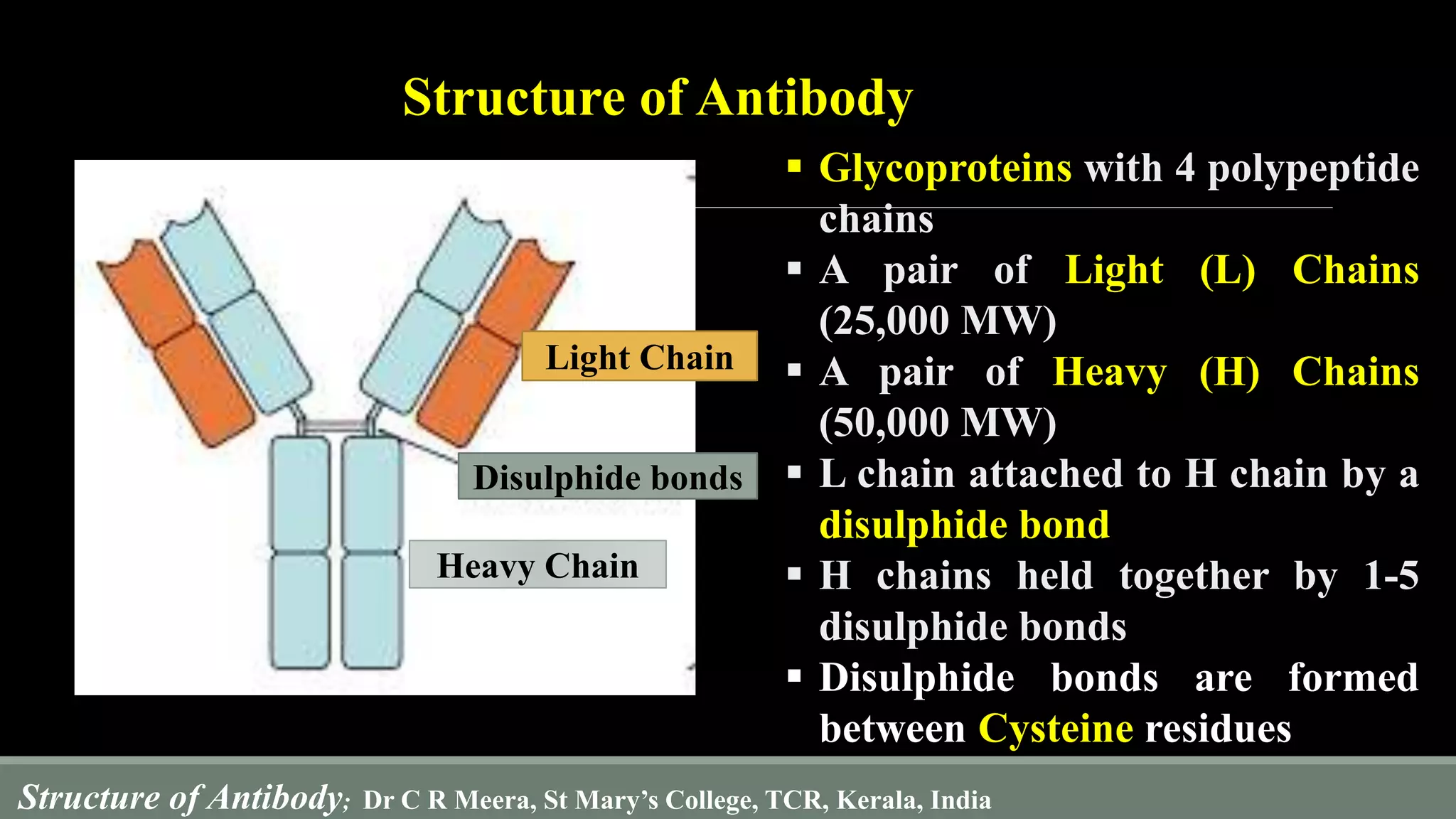
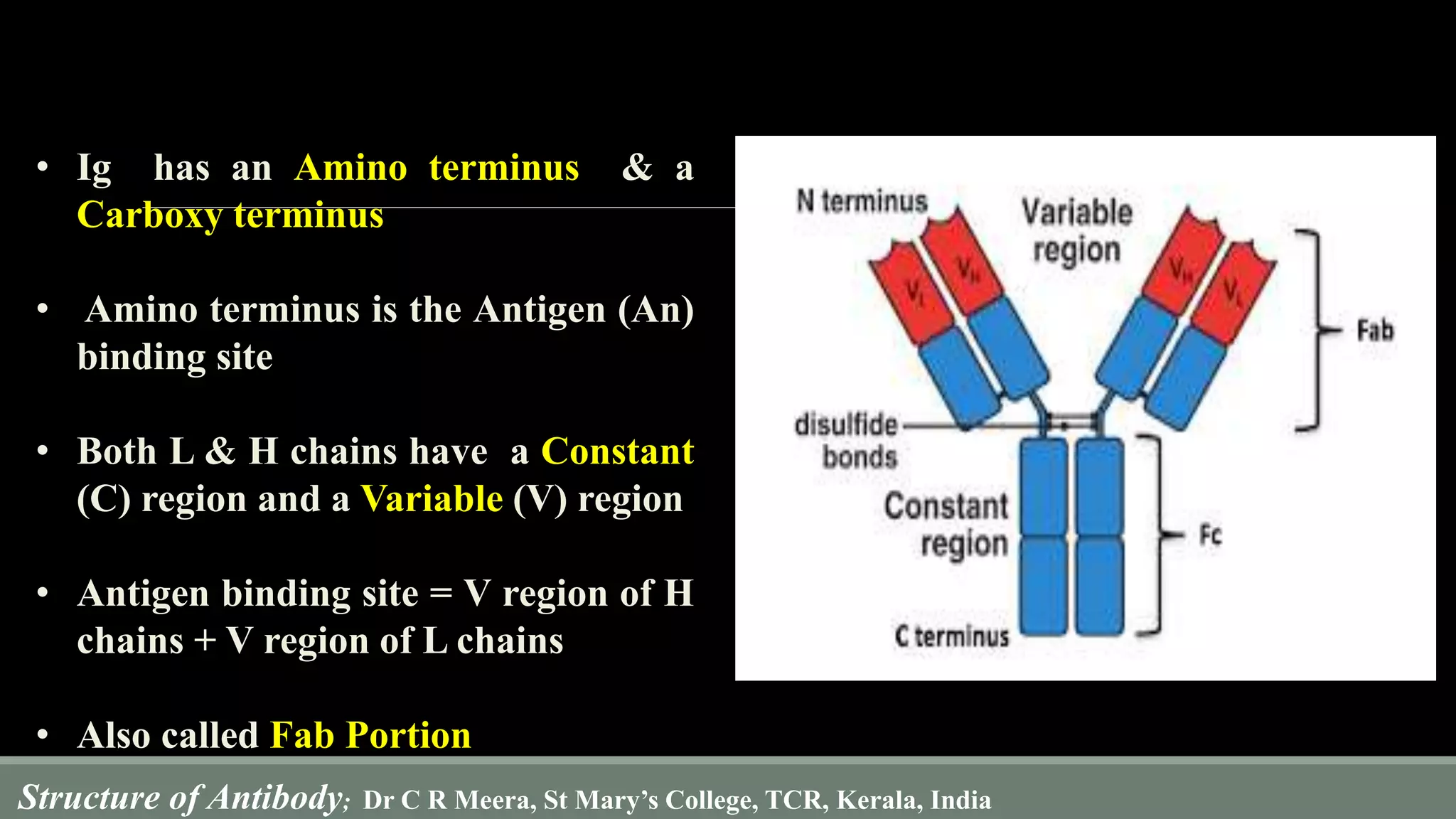
The document describes the structure and function of antibodies, which are proteins produced by B lymphocytes that recognize and neutralize harmful microbes and toxins. It details the components of antibodies, including light and heavy chains, their variable and constant regions, and the types of bonds that stabilize their structure. Additionally, it highlights the differences among the five classes of antibodies and discusses key features such as hypervariable regions and the hinge region's flexibility.