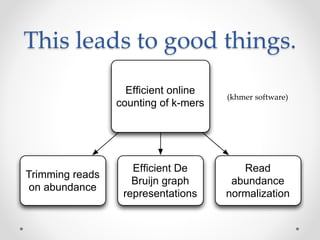
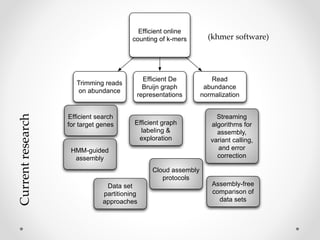
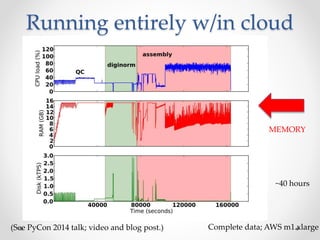
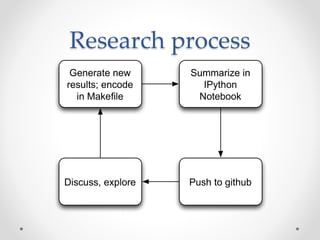
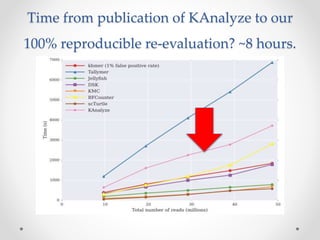
This document discusses openness and reproducibility in computational science. It begins with an introduction and background on the challenges of analyzing non-model organisms. It then describes the goals and challenges of shotgun sequencing analysis, including assembly, counting, and variant calling. It emphasizes the need for efficient data structures, algorithms, and cloud-based analysis to handle large datasets. The document advocates for open science practices like publishing code, data, and analyses to ensure reproducibility of computational results.