This document outlines Dr. Brown's research interests which focus on developing computational tools and approaches to analyze large and complex biological data sets from non-model organisms. Some key points include:
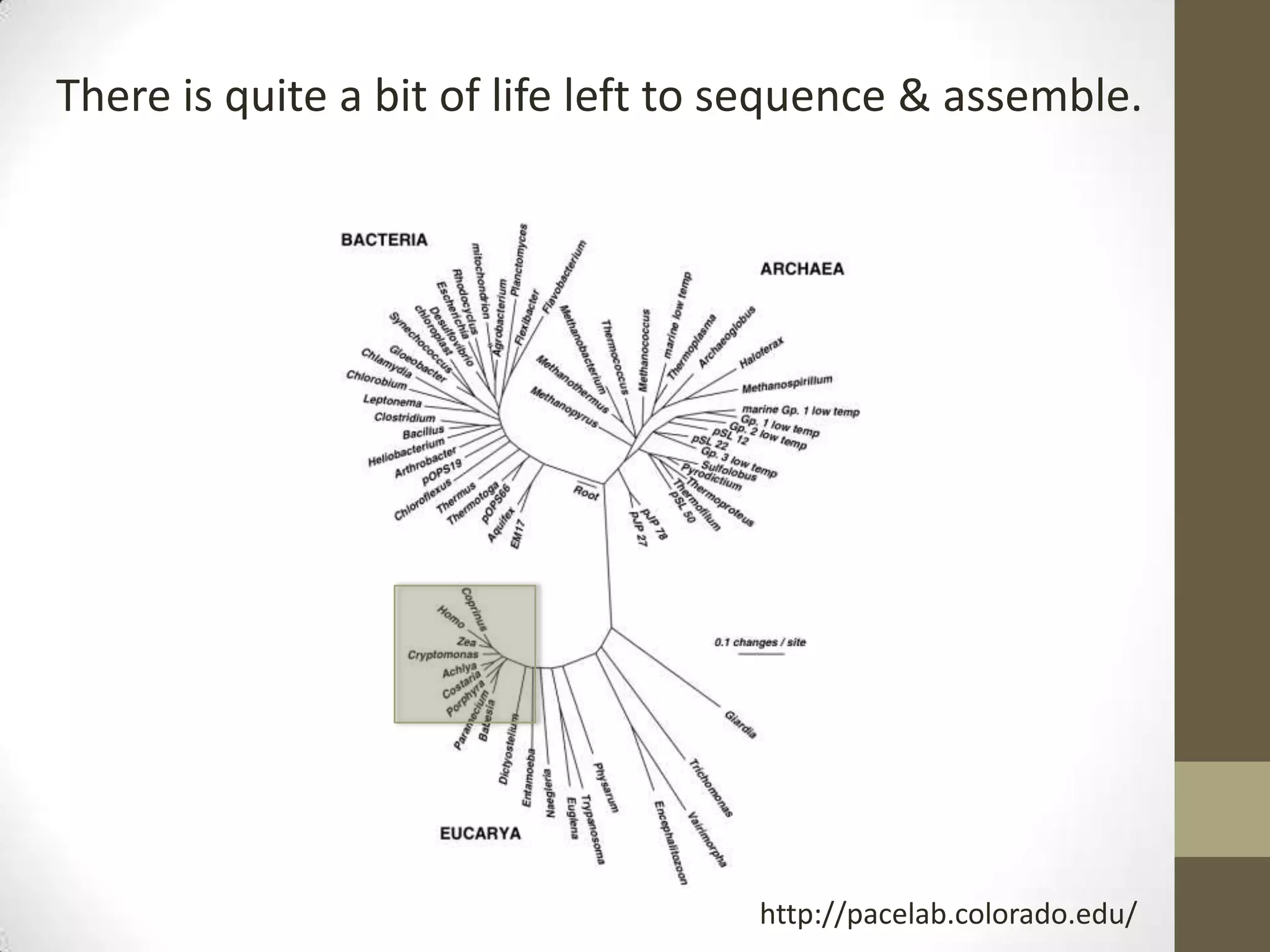
- His lab works on improving genome and transcriptome assembly from samples that are difficult to sequence such as metagenomes, single cells, and samples with high polymorphism.
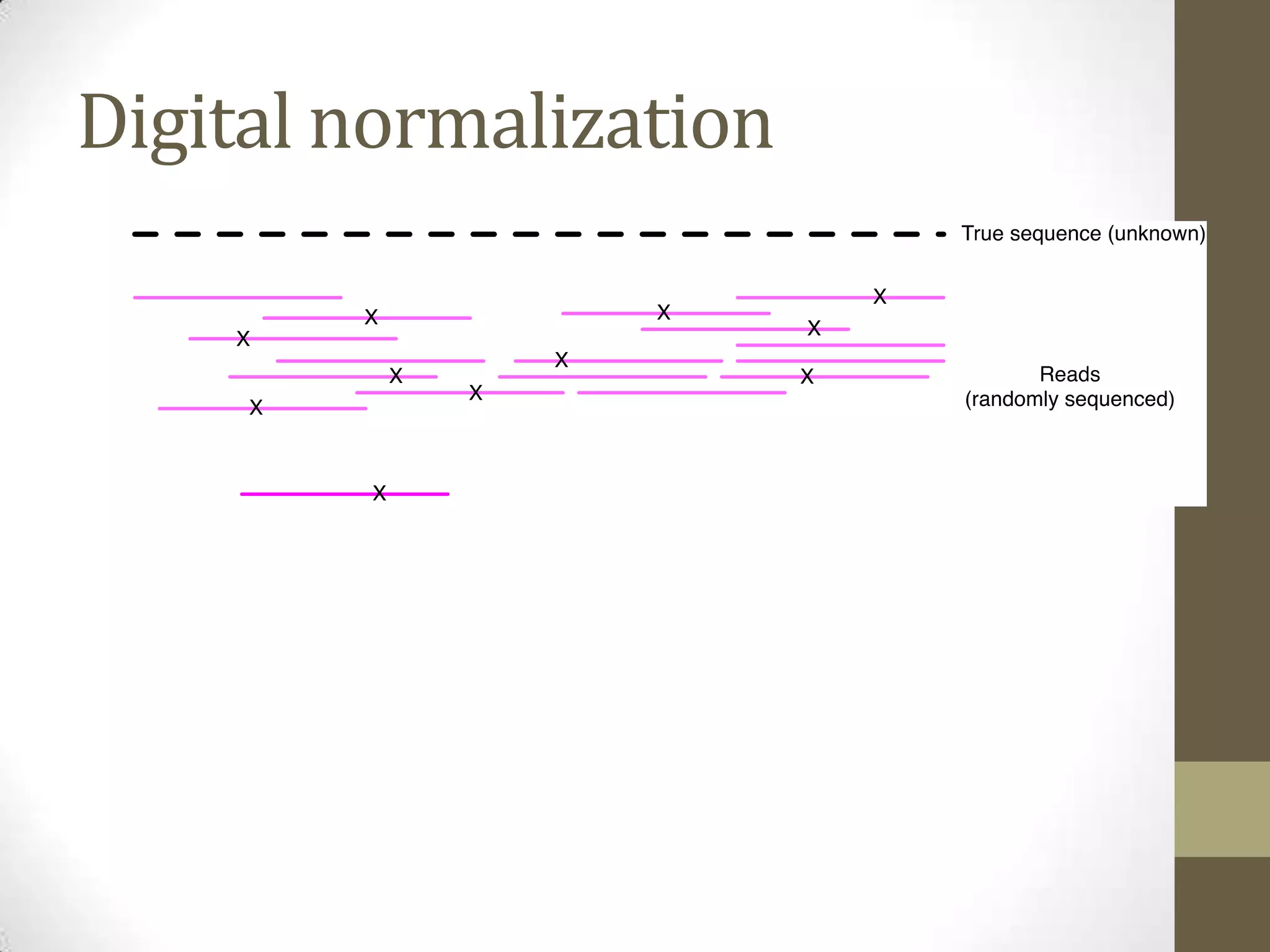
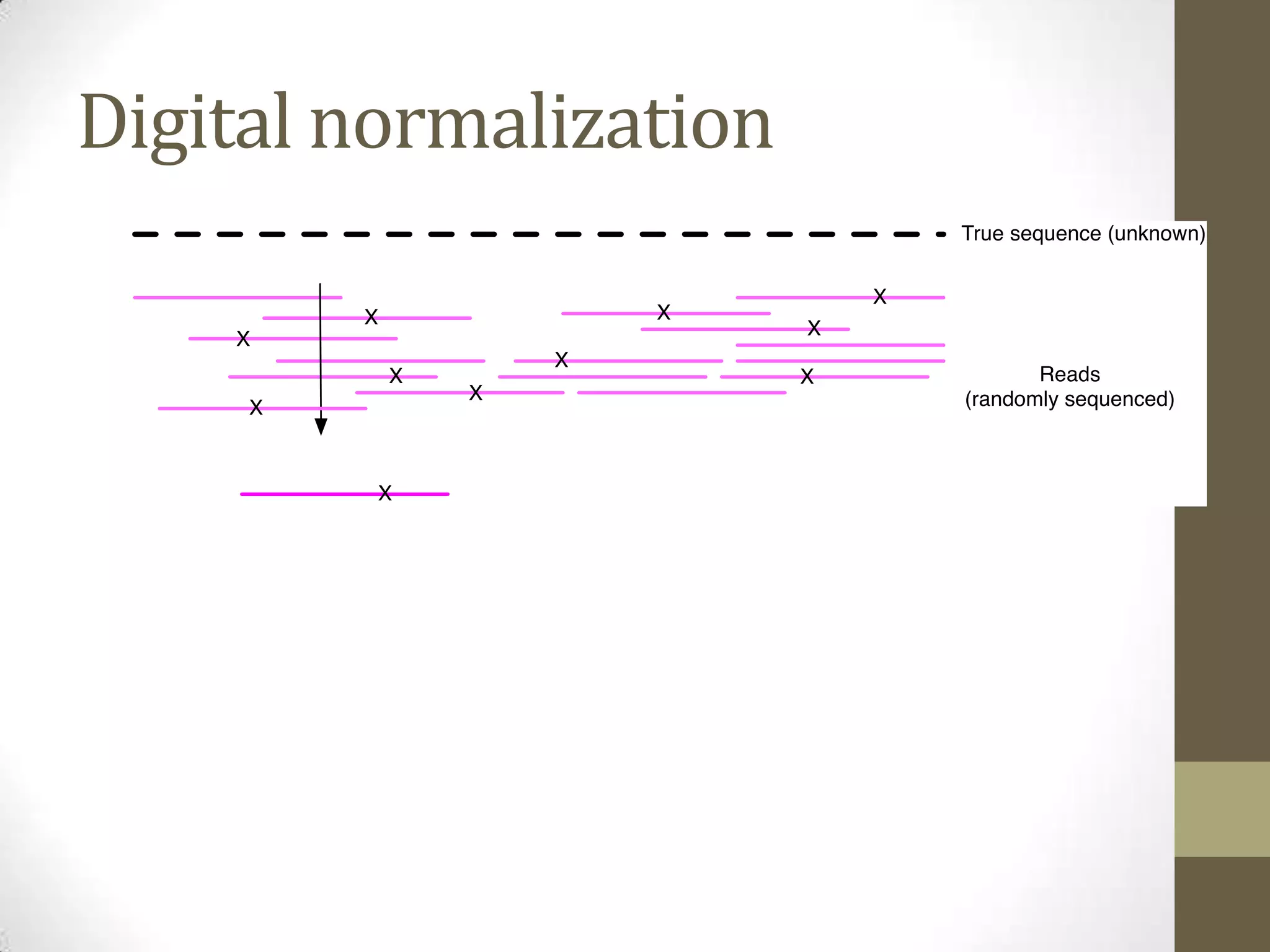
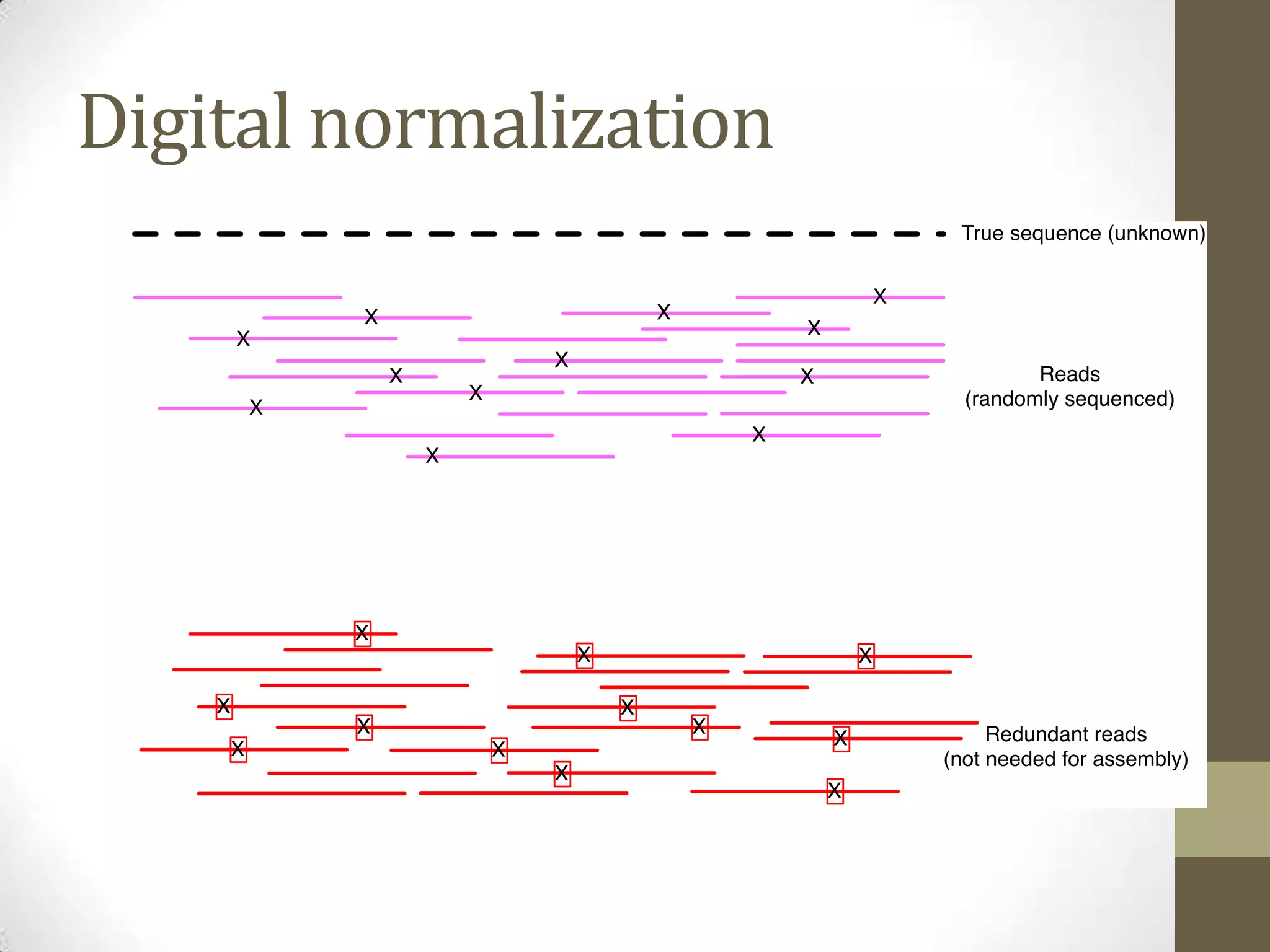
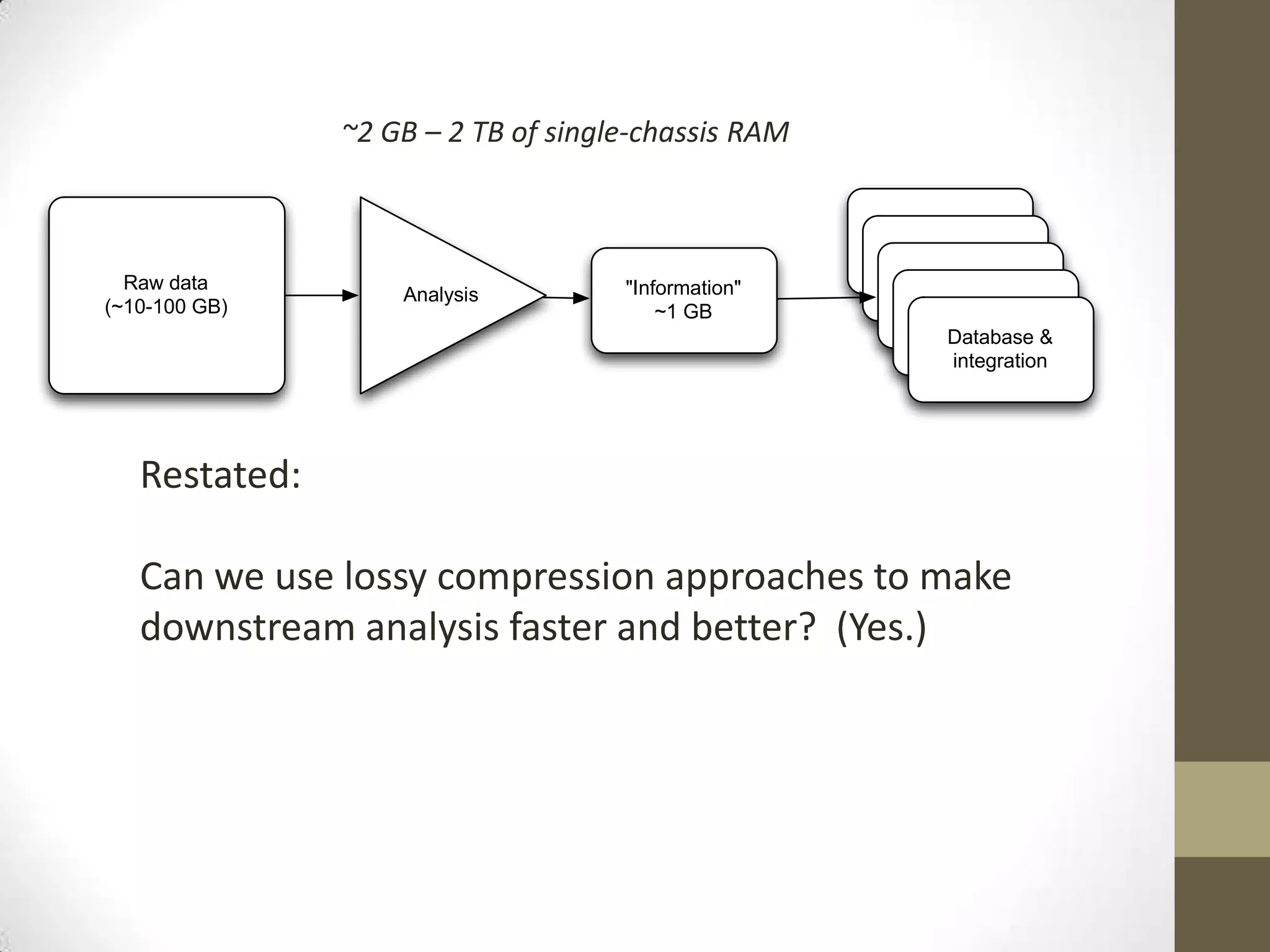
- He describes developing techniques like "digital normalization" which can speed up downstream data analysis by compressing raw data in a lossy but information-preserving way.
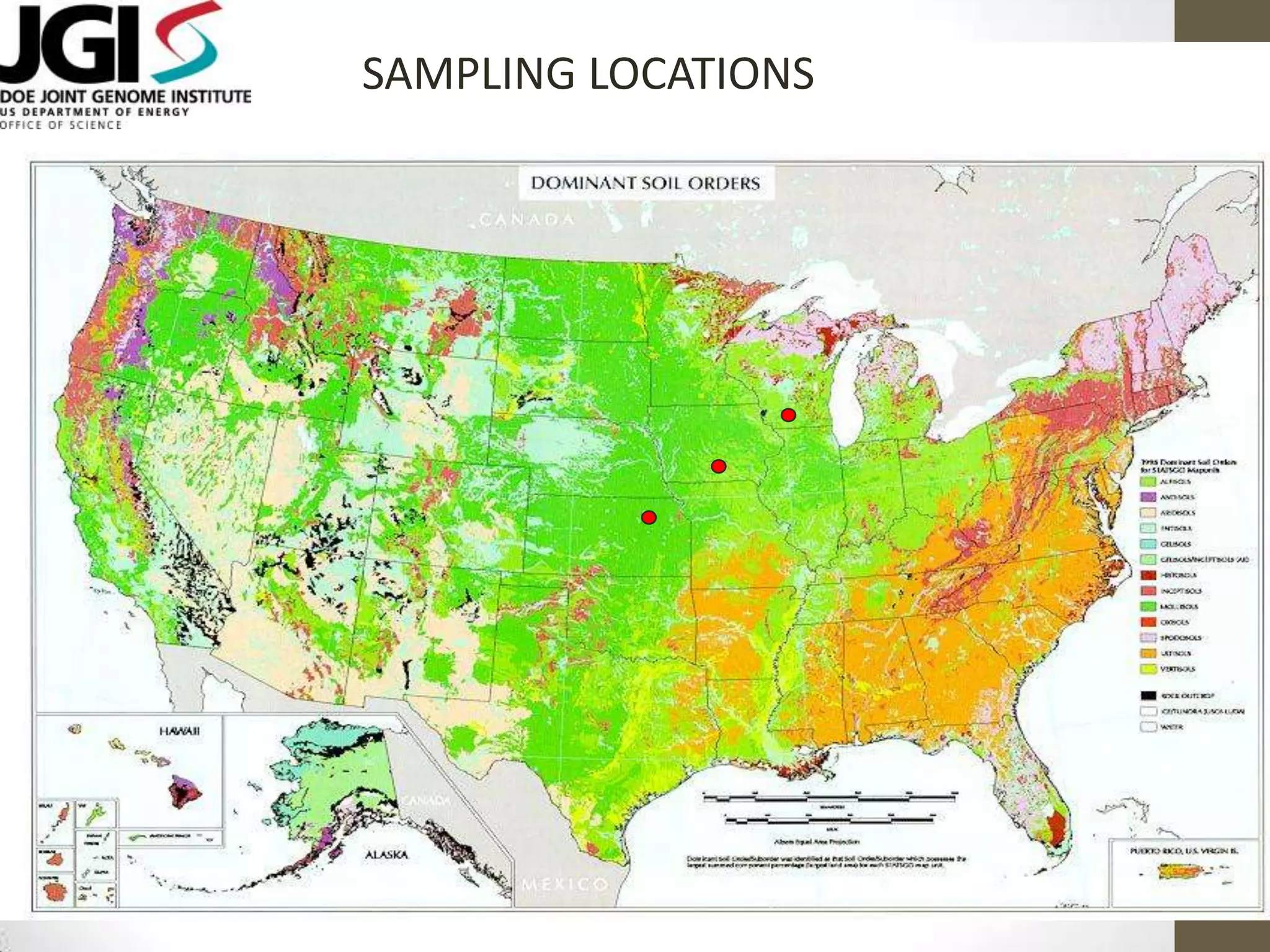
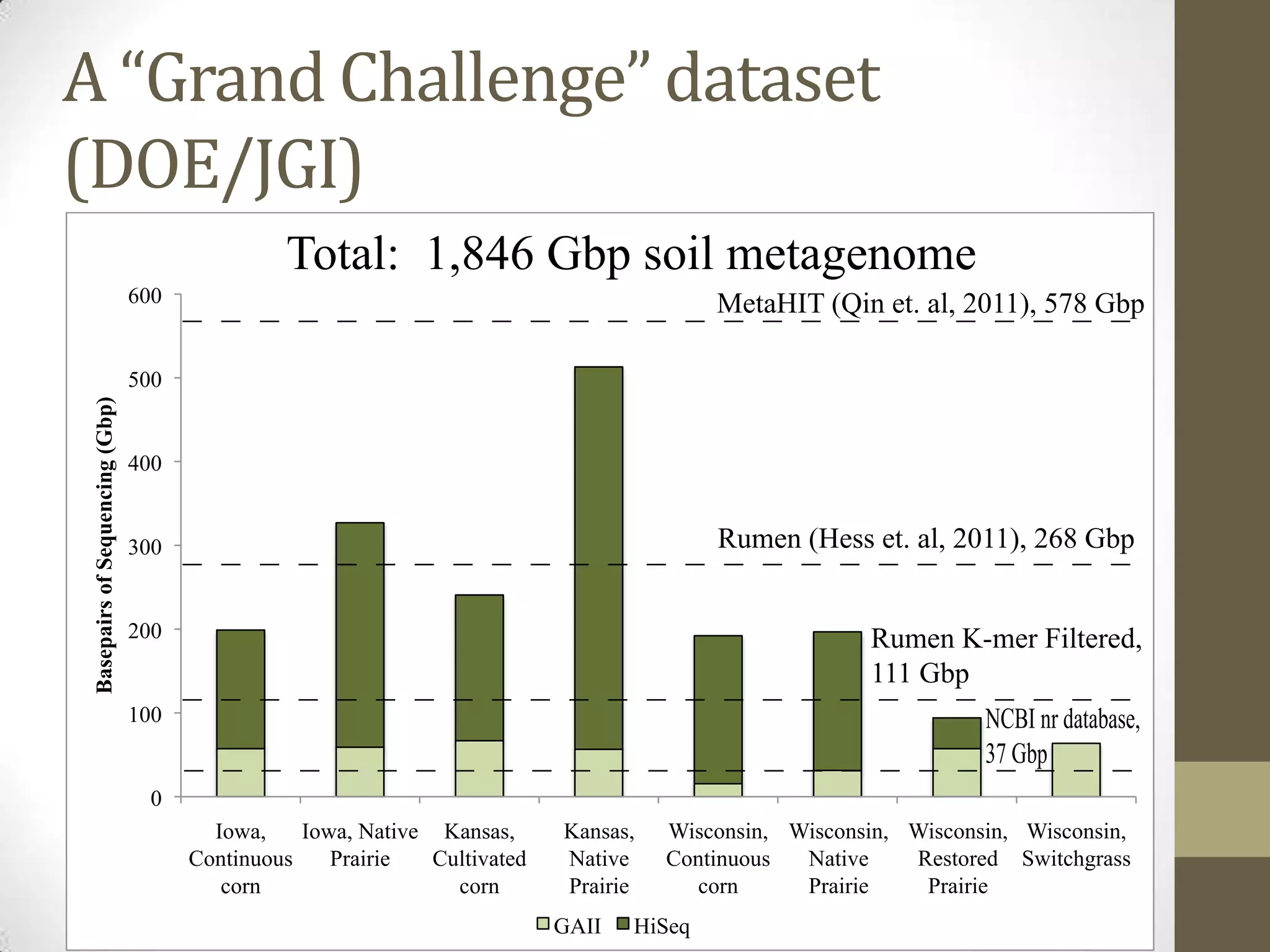
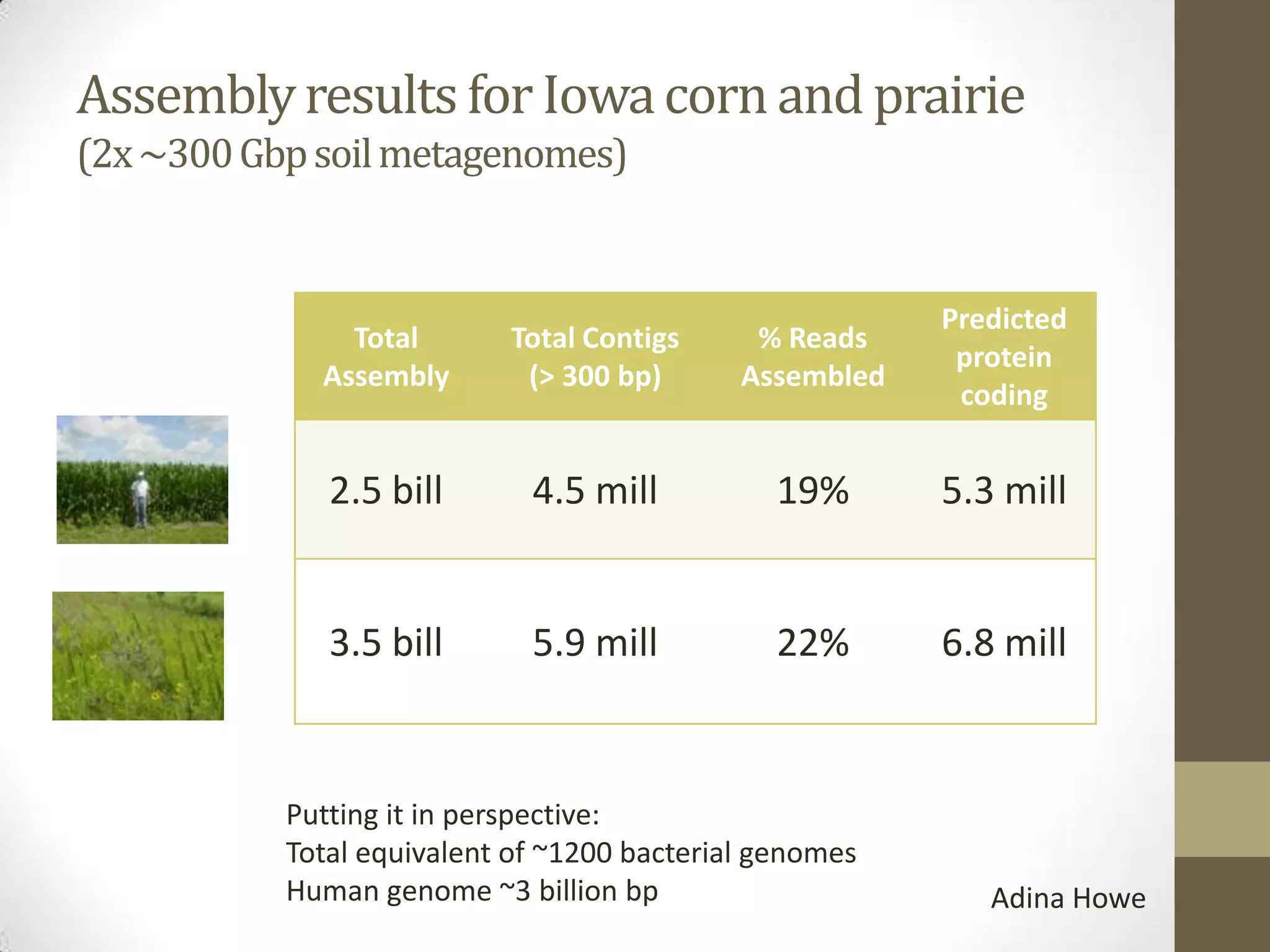
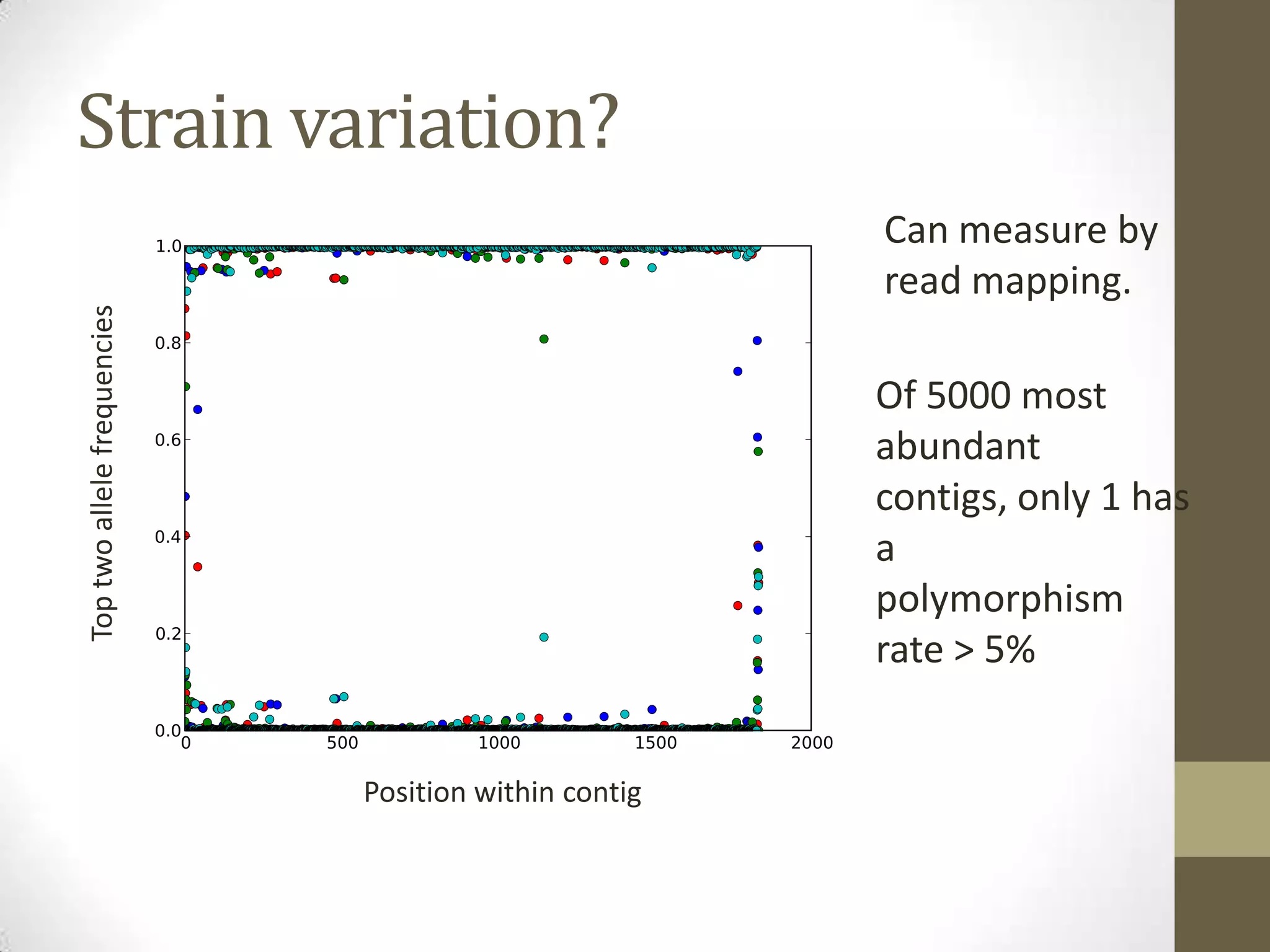
- An example project involved assembling 1.8 terabases of soil metagenome sequence data from various locations, finding little strain-level variation but that phage sequences were abundant