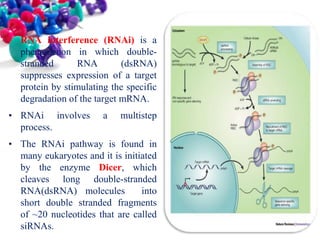
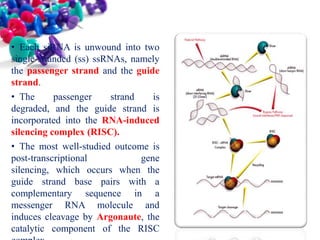
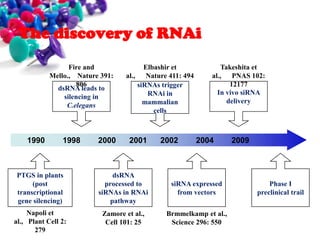
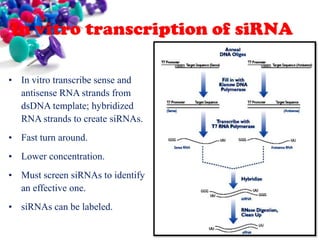
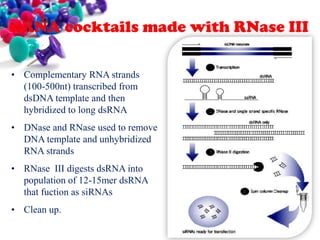
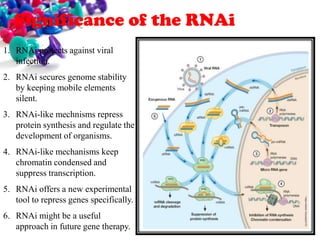
This document discusses small interfering RNA (siRNA), which are double stranded RNA molecules that play a role in RNA interference (RNAi) pathways by interfering with gene expression of complementary nucleotide sequences. siRNAs are naturally produced by the enzyme Dicer but can also be artificially introduced. The document provides details on siRNA structure, the RNAi mechanism, guidelines for effective siRNA design, methods of siRNA synthesis, delivery methods, and applications in gene silencing research and potential therapies.