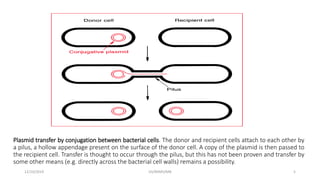
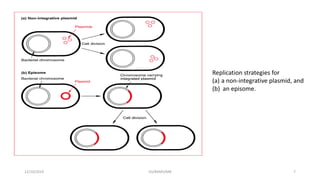
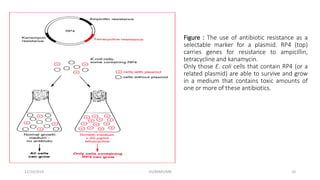
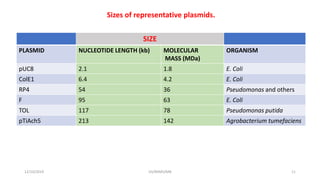
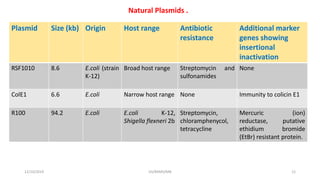
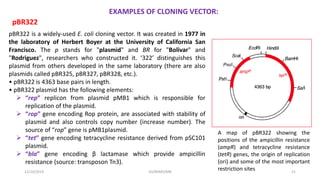
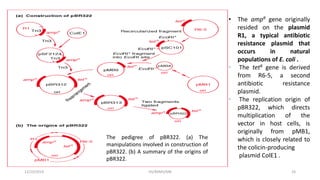
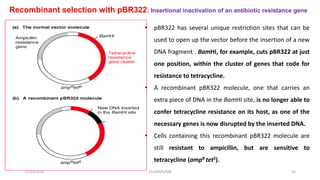
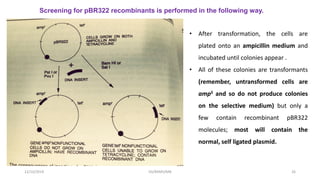
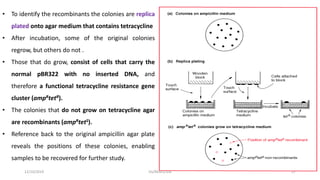
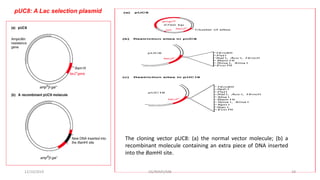
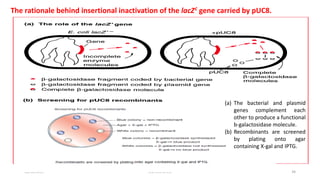
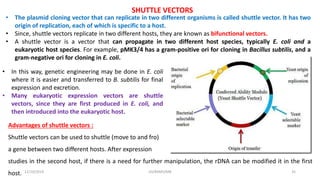
Plasmids are naturally occurring extrachromosomal DNA molecules that can autonomously replicate in bacterial cells. Plasmids are commonly used as cloning vectors to replicate and express exogenous DNA in a host cell. Ideal plasmid vectors are small in size, present in multiple copies, contain origins of replication and selectable markers like antibiotic resistance genes. Commonly used plasmid vectors include pBR322, which contains genes for ampicillin and tetracycline resistance, and has been widely used for cloning in E. coli. Artificial plasmids are also engineered by combining functional modules from different natural plasmids to improve vector characteristics for cloning and expression applications.