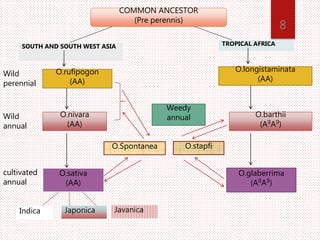
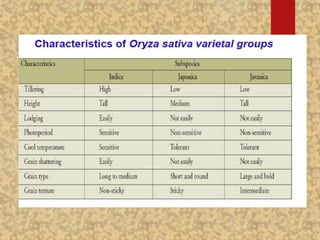
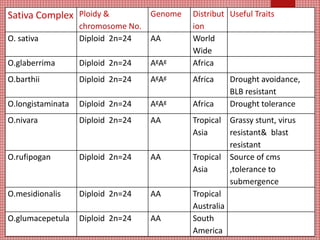
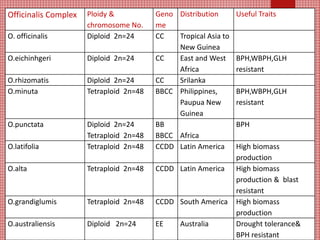
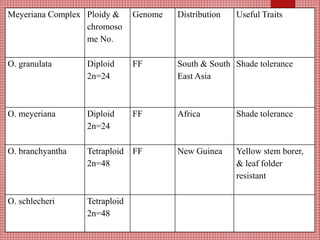
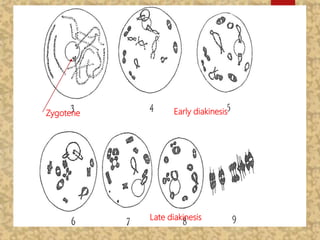
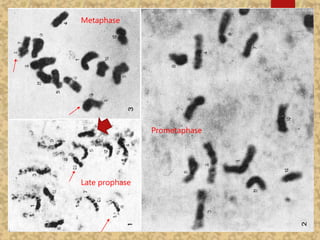
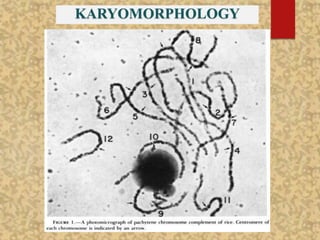
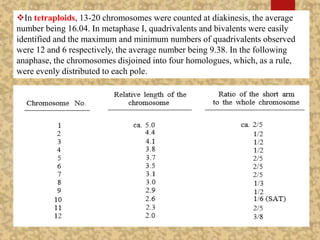
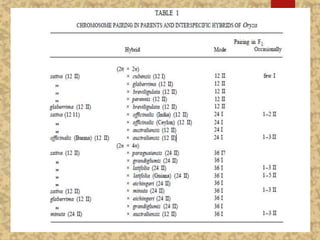
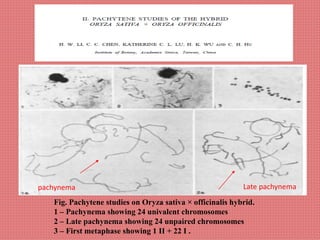
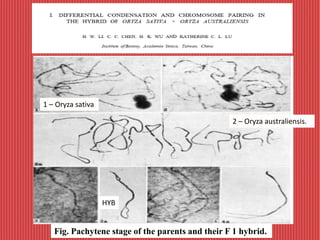
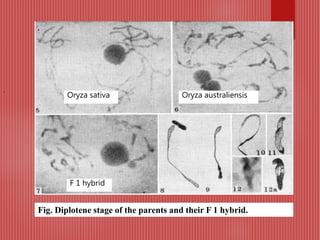
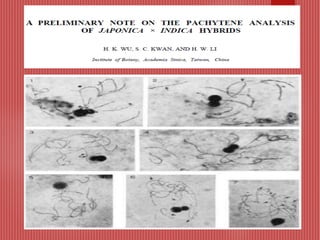
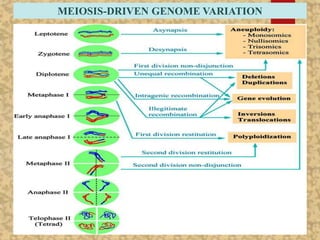
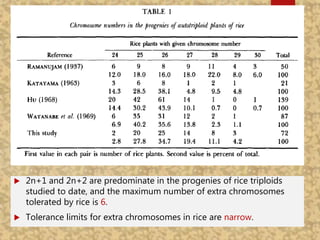
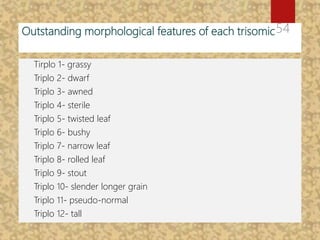
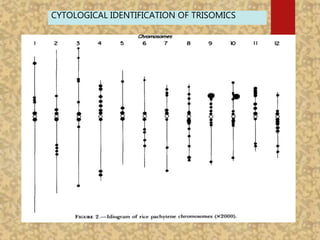
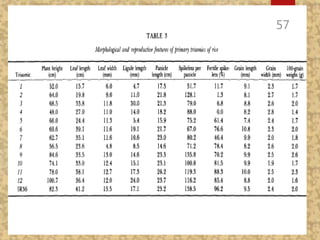
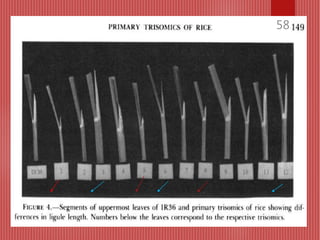
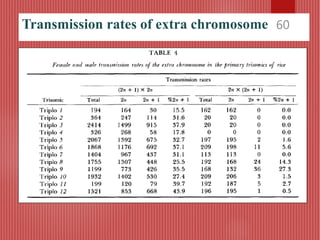
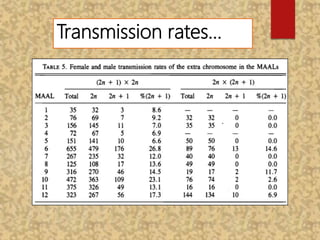
The document discusses the cytogenetics, taxonomy, and karyomorphology of rice, specifically focusing on Oryza sativa and its related species. It explains the origins, chromosome number, structure, and pairing behavior of rice chromosomes, including insights from various researchers over time. The document highlights the genetic diversity among rice species and their economic significance, particularly in terms of beneficial traits from wild relatives.