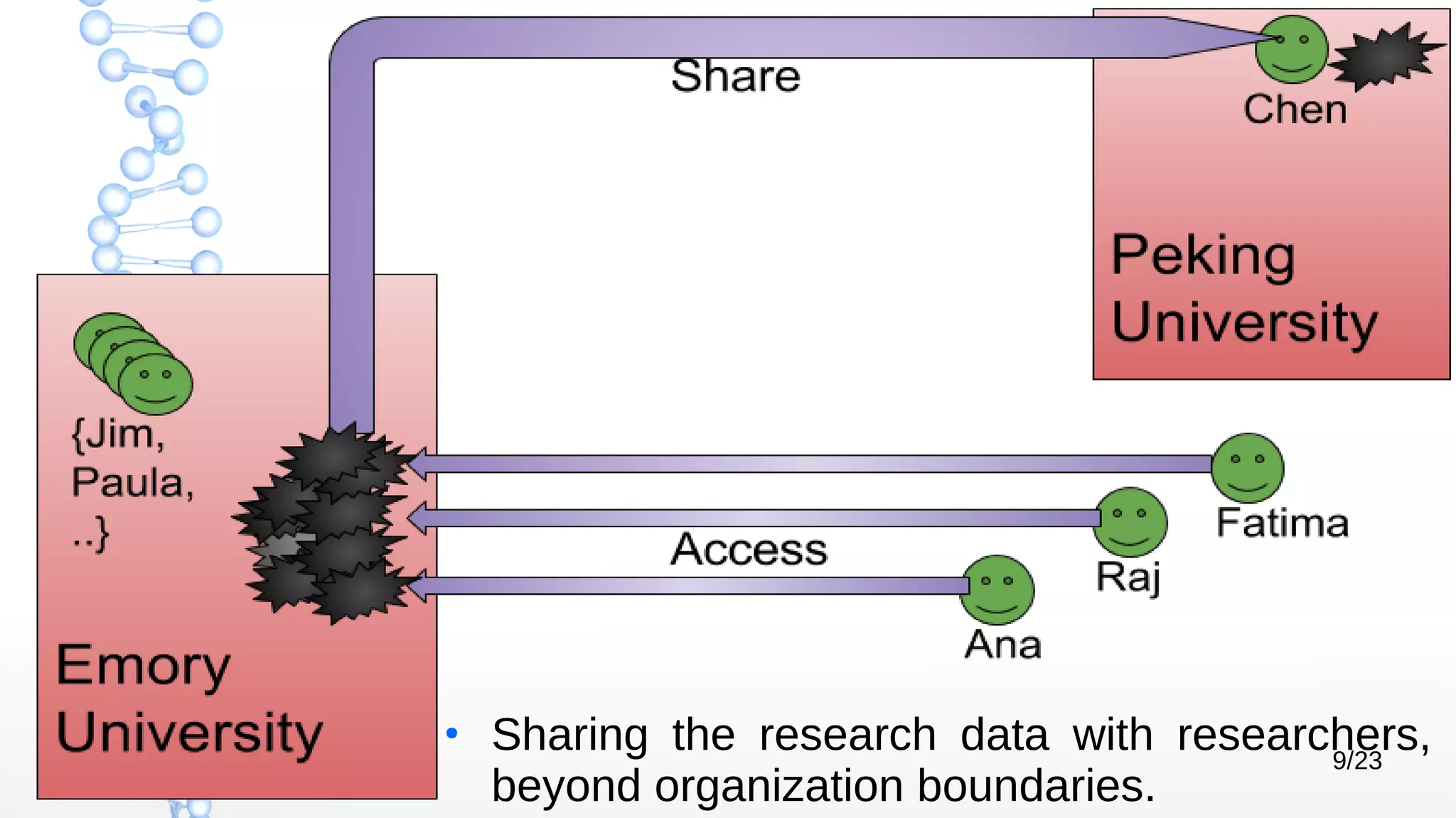
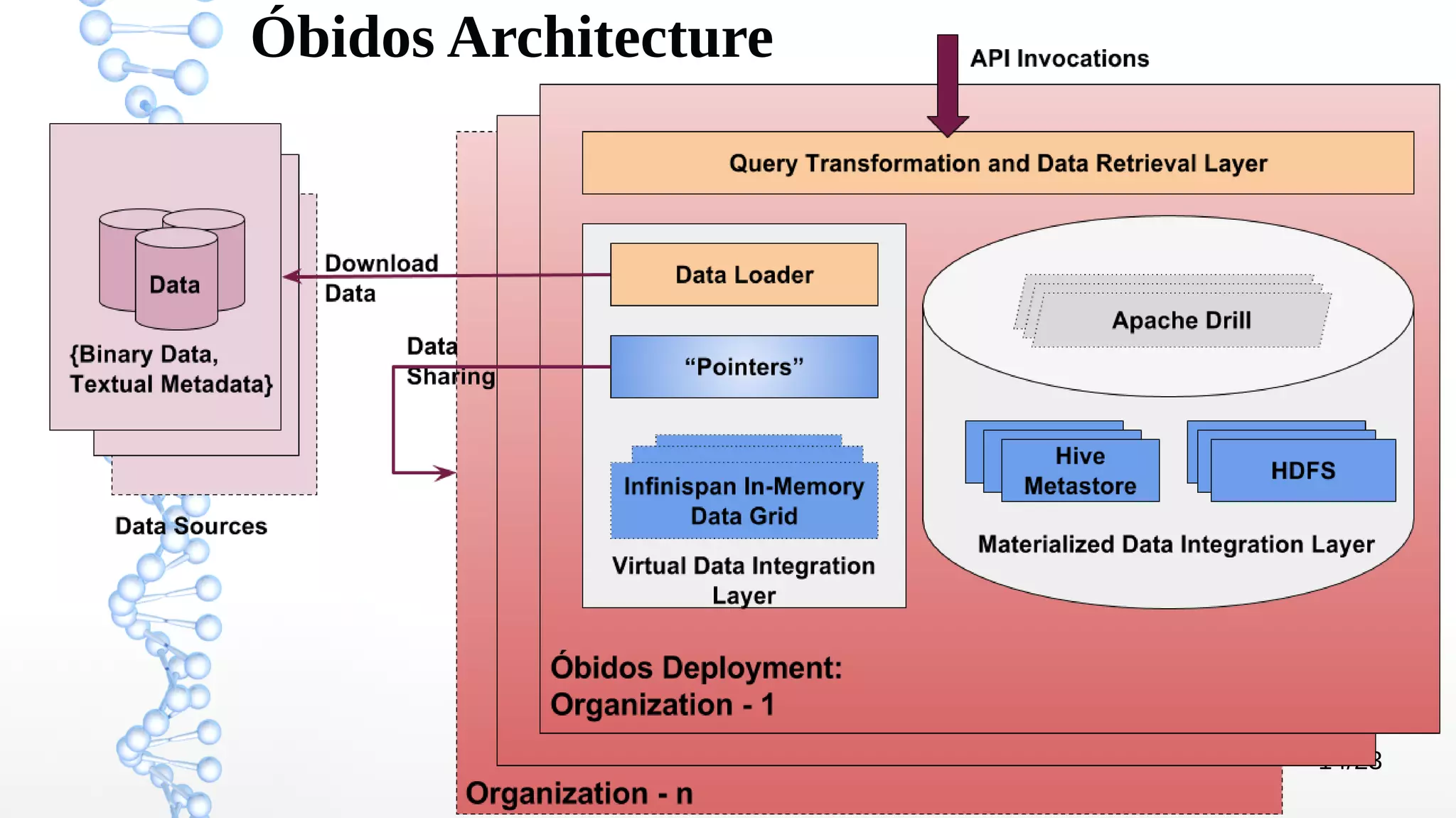
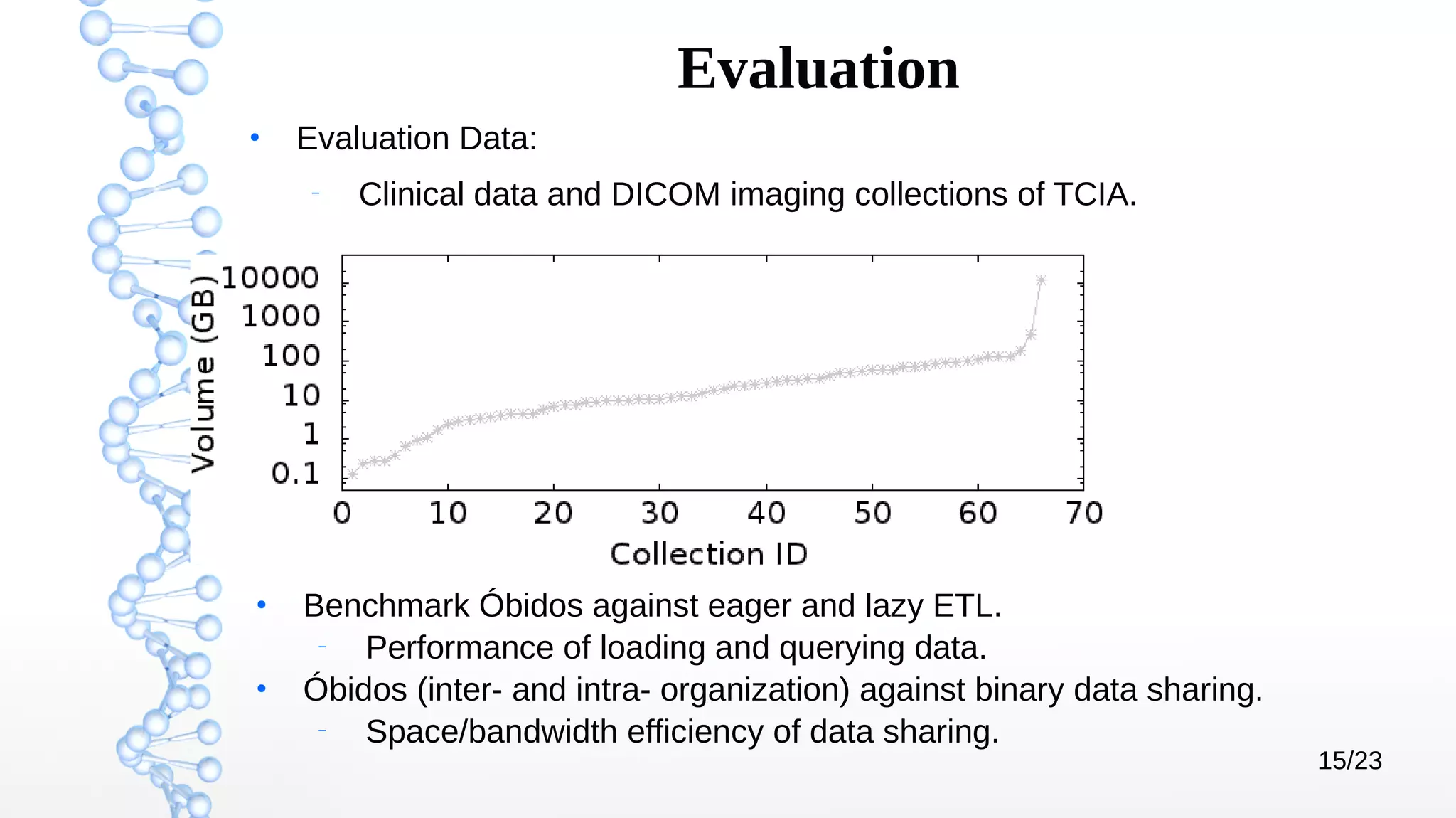
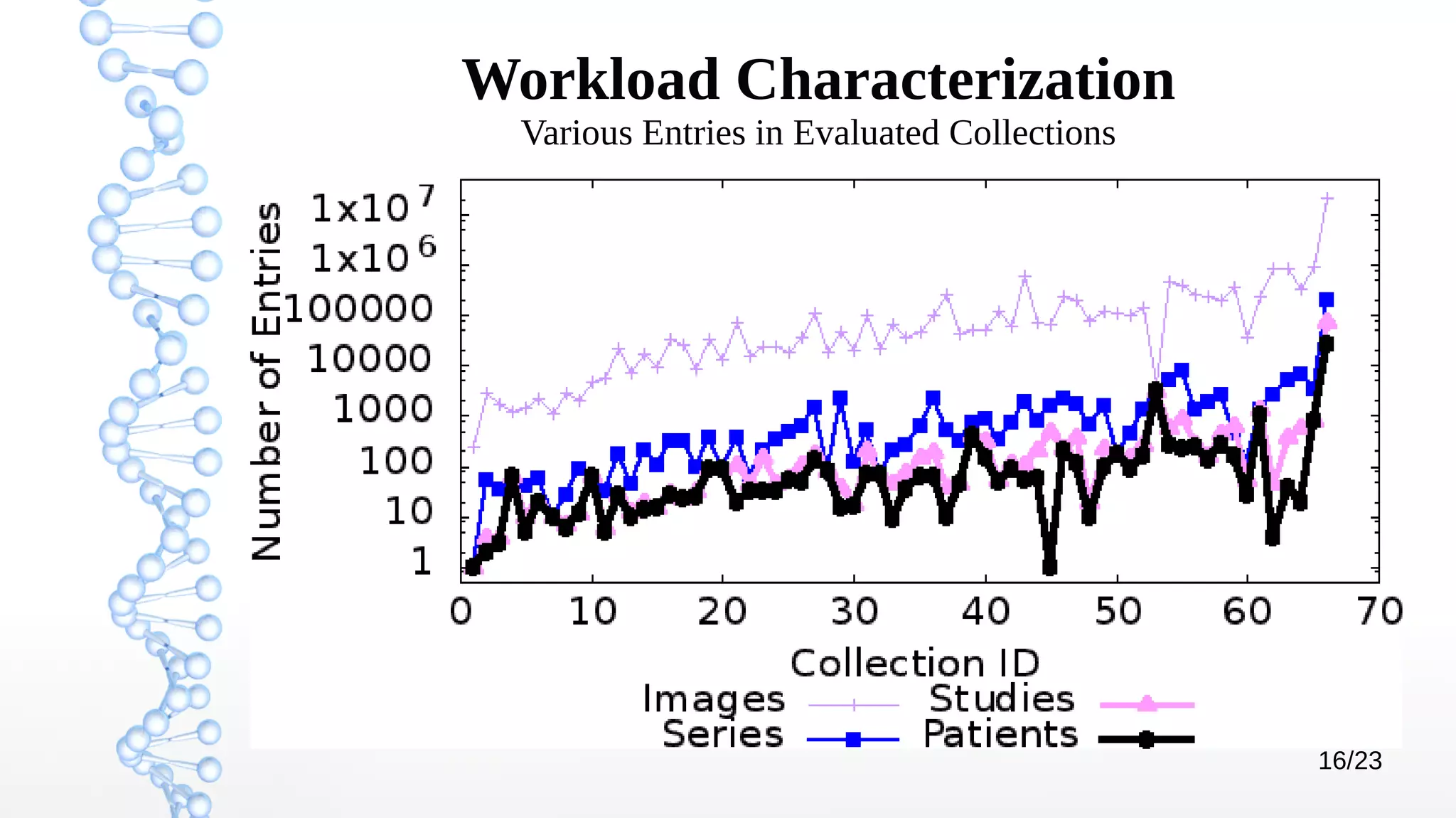
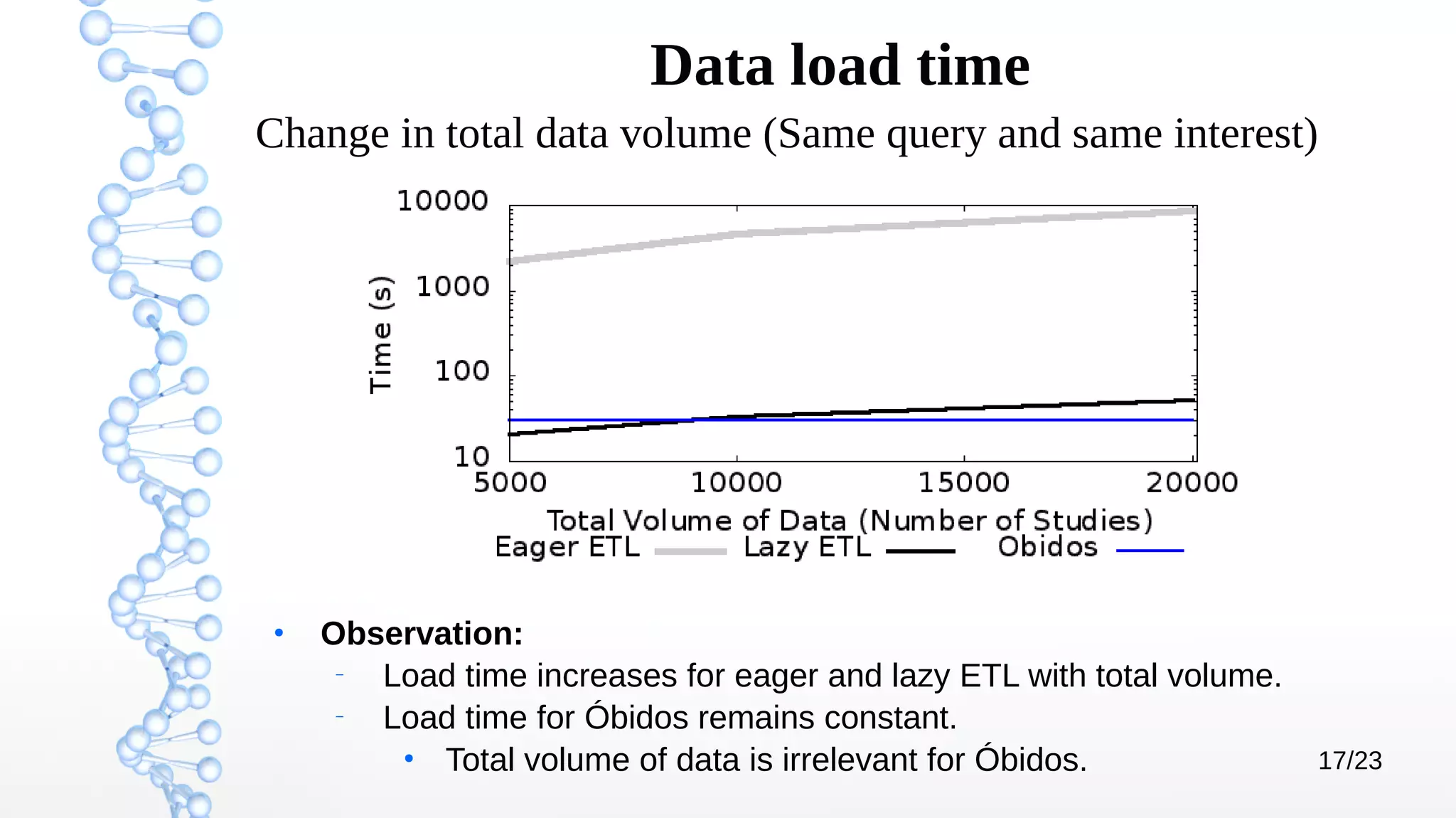
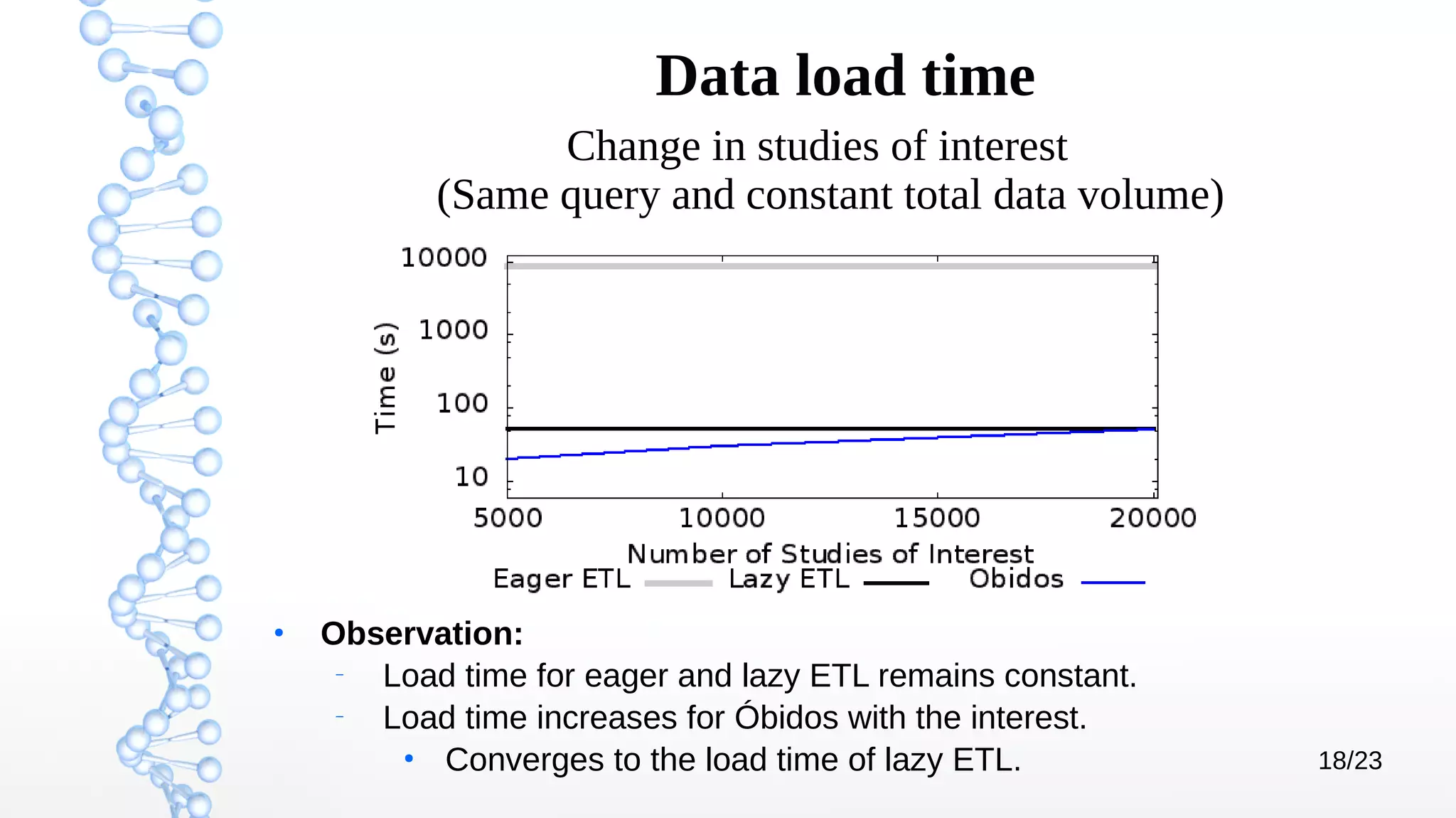
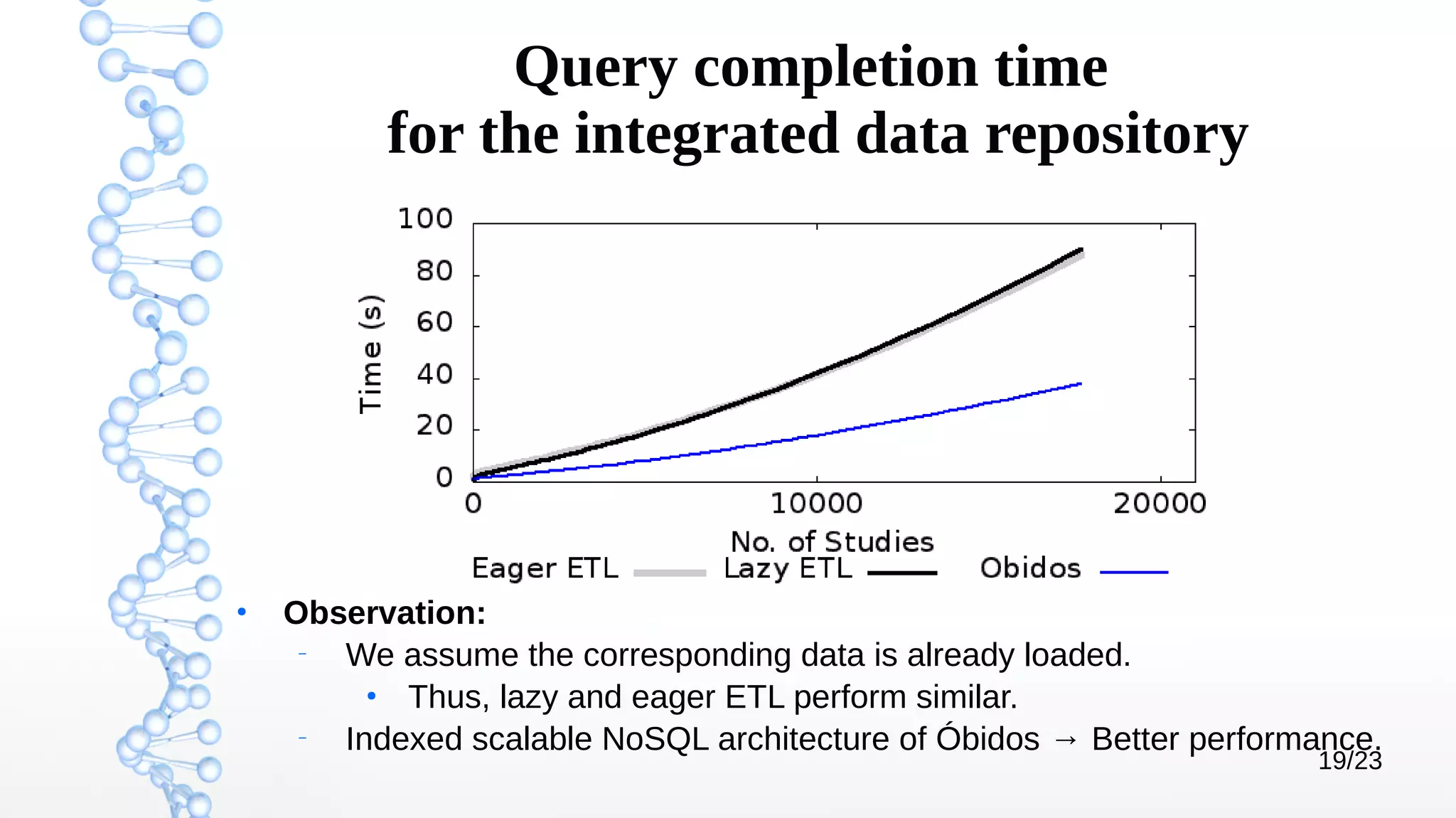
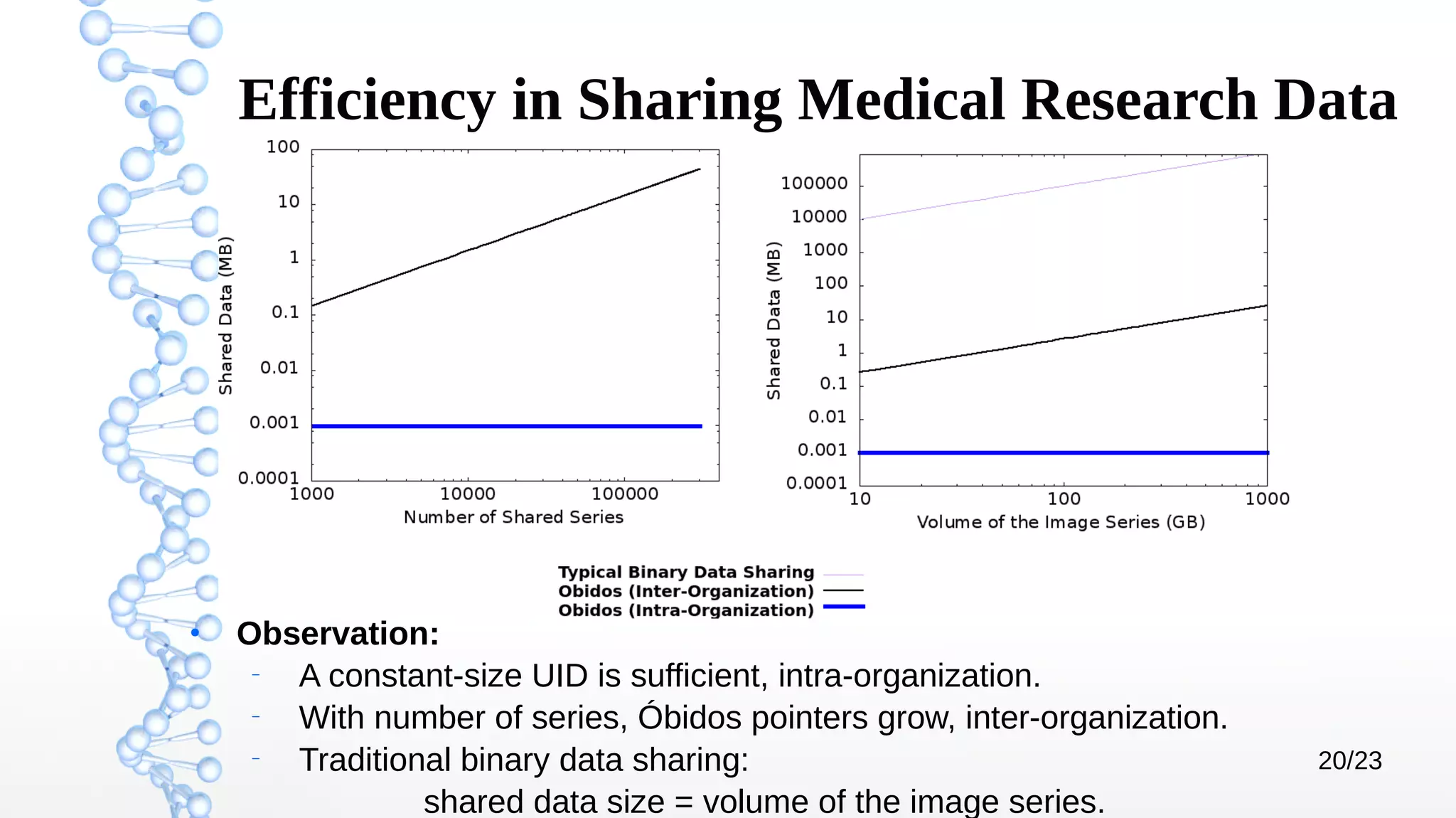
The document presents Óbidos, a system designed for on-demand service-based big data integration specifically optimized for medical research collaboration. It addresses various challenges in integrating diverse data types and sources, aiming to enhance efficiency in data access and sharing while avoiding data duplication. The evaluation indicates that Óbidos provides fast, efficient data analysis and effective sharing mechanisms compared to traditional binary data sharing methods.